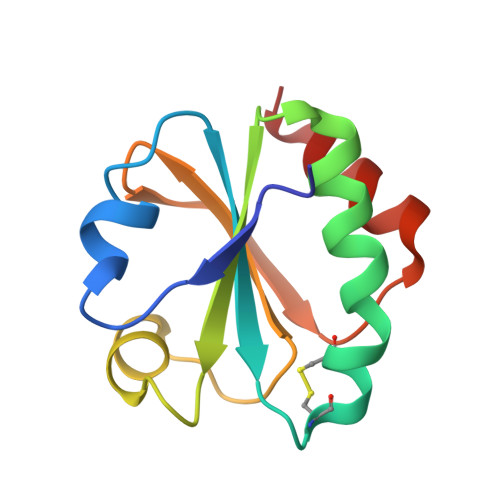
Atomic-resolution crystal structure of thioredoxin from the acidophilic bacterium Acetobacter aceti.
Starks, C.M., Francois, J.A., Macarthur, K.M., Heard, B.Z., Kappock, T.J.(2007) Protein Sci 16: 92-98
- PubMed: 17192591 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.062519707
- Primary Citation Related Structures:
2I4A - PubMed Abstract:
The crystal structure of thioredoxin (AaTrx) from the acetic acid bacterium Acetobacter aceti was determined at 1 A resolution. This is currently the highest resolution crystal structure available for any thioredoxin. Thioredoxins facilitate thiol-disulfide exchange, a process that is expected to be slow at the low pH values encountered in the A. aceti cytoplasm. Despite the apparent need to function at low pH, neither the active site nor the surface charge distribution of AaTrx is notably different from that of Escherichia coli thioredoxin. Apparently the ancestral thioredoxin was sufficiently stable for use in A. aceti or the need to interact with multiple targets constrained the variation of surface residues. The AaTrx structure presented here provides a clear view of all ionizable protein moieties and waters, a first step in understanding how thiol-disulfide exchange might occur in a low pH cytoplasm, and is a basis for biophysical studies of the mechanism of acid-mediated unfolding. The high resolution of this structure should be useful for computational studies of thioredoxin function, protein structure and dynamics, and side-chain ionization.
- Department of Chemistry, Washington University in Saint Louis, Missouri 63130, USA.
Organizational Affiliation: