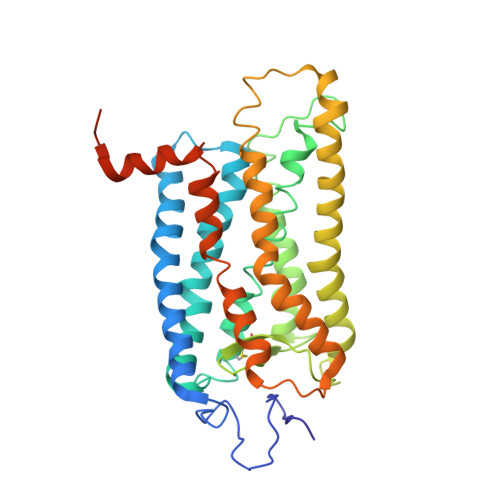
Crystal structure of a photoactivated deprotonated intermediate of rhodopsin.
Salom, D., Lodowski, D.T., Stenkamp, R.E., Trong, I.L., Golczak, M., Jastrzebska, B., Harris, T., Ballesteros, J.A., Palczewski, K.(2006) Proc Natl Acad Sci U S A 103: 16123-16128
- PubMed: 17060607 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0608022103
- Primary Citation Related Structures:
2I35, 2I36, 2I37 - PubMed Abstract:
The changes that lead to activation of G protein-coupled receptors have not been elucidated at the structural level. In this work we report the crystal structures of both ground state and a photoactivated deprotonated intermediate of bovine rhodopsin at a resolution of 4.15 A. In the photoactivated state, the Schiff base linking the chromophore and Lys-296 becomes deprotonated, reminiscent of the G protein-activating state, metarhodopsin II. The structures reveal that the changes that accompany photoactivation are smaller than previously predicted for the metarhodopsin II state and include changes on the cytoplasmic surface of rhodopsin that possibly enable the coupling to its cognate G protein, transducin. Furthermore, rhodopsin forms a potentially physiologically relevant dimer interface that involves helices I, II, and 8, and when taken with the prior work that implicates helices IV and V as the physiological dimer interface may account for one of the interfaces of the oligomeric structure of rhodopsin seen in the membrane by atomic force microscopy. The activation and oligomerization models likely extend to the majority of other G protein-coupled receptors.
- Novasite Pharmaceuticals Inc., San Diego, CA 92121, USA.
Organizational Affiliation: