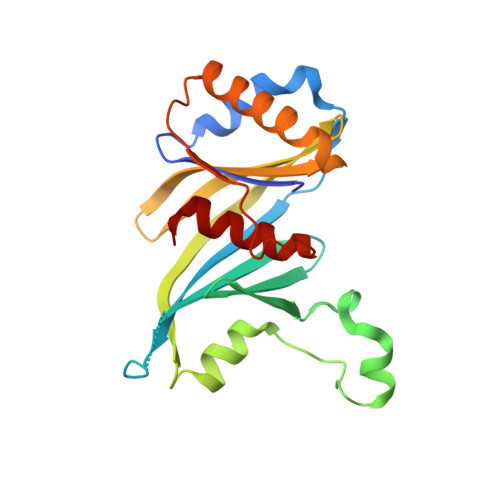
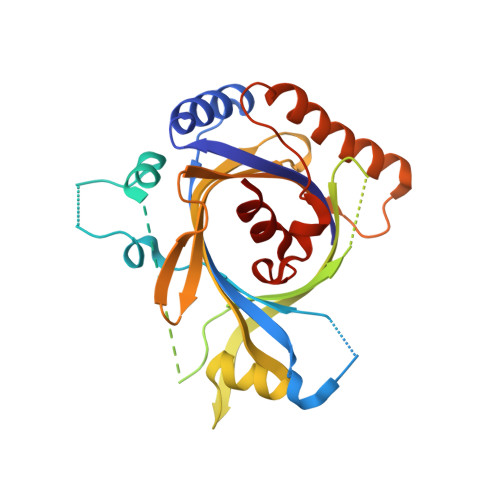
Structure and TBP binding of the Mediator head subcomplex Med8-Med18-Med20.
Lariviere, L., Geiger, S., Hoeppner, S., Rother, S., Strasser, K., Cramer, P.(2006) Nat Struct Mol Biol 13: 895-901
- PubMed: 16964259 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb1143
- Primary Citation Related Structures:
2HZM, 2HZS - PubMed Abstract:
The Mediator head module stimulates basal RNA polymerase II (Pol II) transcription and enables transcriptional regulation. Here we show that the head subunits Med8, Med18 and Med20 form a subcomplex (Med8/18/20) with two submodules. The highly conserved N-terminal domain of Med8 forms one submodule that binds the TATA box-binding protein (TBP) in vitro and is essential in vivo. The second submodule consists of the C-terminal region of Med8 (Med8C), Med18 and Med20. X-ray analysis of this submodule reveals that Med18 and Med20 form related beta-barrel folds. A conserved putative protein-interaction face on the Med8C/18/20 submodule includes sites altered by srb mutations, which counteract defects resulting from Pol II truncation. Our results and published data support a positive role of the Med8/18/20 subcomplex in initiation-complex formation and suggest that the Mediator head contains a multipartite TBP-binding site that can be modulated by transcriptional activators.
- Gene Center Munich, Department of Chemistry and Biochemistry, Ludwig-Maximilians-Universität München, Feodor-Lynen-Str. 25, 81377 Munich, Germany.
Organizational Affiliation: