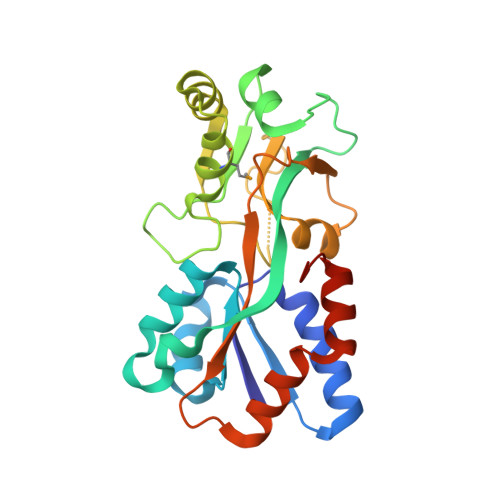
Structural context for protein N-glycosylation in bacteria: The structure of PEB3, an adhesin from Campylobacter jejuni.
Rangarajan, E.S., Bhatia, S., Watson, D.C., Munger, C., Cygler, M., Matte, A., Young, N.M.(2007) Protein Sci 16: 990-995
- PubMed: 17456748 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.062737507
- Primary Citation Related Structures:
2HXW - PubMed Abstract:
Campylobacter jejuni is unusual among bacteria in possessing a eukaryotic-like system for N-linked protein glycosylation at Asn residues in sequons of the type Asp/Glu-Xaa-Asn-Xaa-Ser/Thr. However, little is known about the structural context of the glycosylated sequons, limiting the design of novel recombinant glycoproteins. To obtain more information on sequon structure, we have determined the crystal structure of the PEB3 (Cj0289c) dimer. PEB3 has the class II periplasmic-binding protein fold, with each monomer having two domains with a ligand-binding site containing citrate located between them, and overall resembles molybdate- and sulfate-binding proteins. The sequon around Asn90 is located within a surface-exposed loop joining two structural elements. The three key residues are well exposed on the surface; hence, they may be accessible to the PglB oligosaccharyltransferase in the folded state.
- Department of Biochemistry, McGill University, Montreal, Quebec H3G 1Y6, Canada.
Organizational Affiliation: