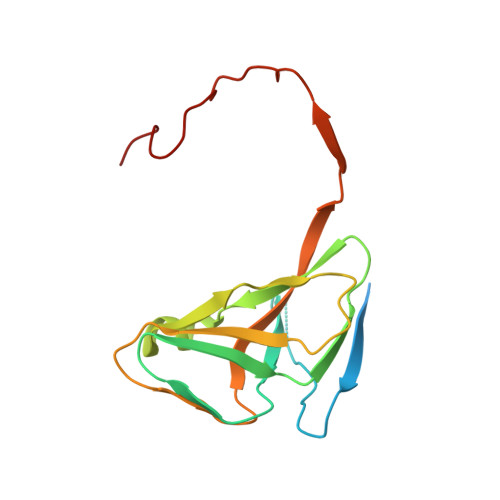
Active site closure facilitates juxtaposition of reactant atoms for initiation of catalysis by human dUTPase.
Varga, B., Barabas, O., Kovari, J., Toth, J., Hunyadi-Gulyas, E., Klement, E., Medzihradszky, K.F., Tolgyesi, F., Fidy, J., Vertessy, B.G.(2007) FEBS Lett 581: 4783-4788
- PubMed: 17880943 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2007.09.005
- Primary Citation Related Structures:
2HQU - PubMed Abstract:
Human dUTPase, essential for DNA integrity, is an important survival factor for cancer cells. We determined the crystal structure of the enzyme:alpha,beta-imino-dUTP:Mg complex and performed equilibrium binding experiments in solution. Ordering of the C-terminus upon the active site induces close juxtaposition of the incoming nucleophile attacker water oxygen and the alpha-phosphorus of the substrate, decreasing their distance below the van der Waals limit. Complex interactions of the C-terminus with both substrate and product were observed via a specifically designed tryptophan sensor, suitable for further detailed kinetic and ligand binding studies. Results explain the key functional role of the C-terminus.
- Institute of Enzymology, Biological Research Center, Hungarian Academy of Sciences, Budapest, Hungary.
Organizational Affiliation: