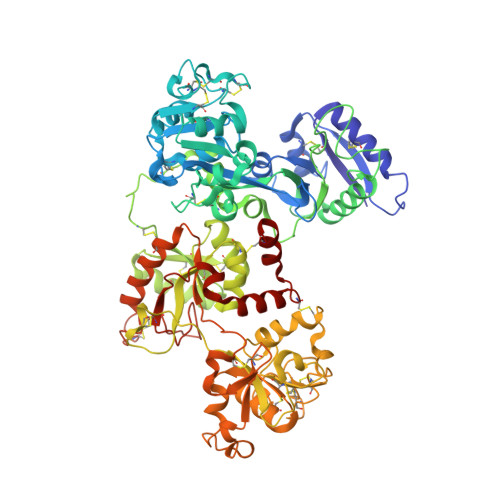
The Crystal Structure of Iron-free Human Serum Transferrin Provides Insight into Inter-lobe Communication and Receptor Binding.
Wally, J., Halbrooks, P.J., Vonrhein, C., Rould, M.A., Everse, S.J., Mason, A.B., Buchanan, S.K.(2006) J Biological Chem 281: 24934-24944
- PubMed: 16793765 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M604592200
- Primary Citation Related Structures:
2HAU, 2HAV - PubMed Abstract:
Serum transferrin reversibly binds iron in each of two lobes and delivers it to cells by a receptor-mediated, pH-dependent process. The binding and release of iron result in a large conformational change in which two subdomains in each lobe close or open with a rigid twisting motion around a hinge. We report the structure of human serum transferrin (hTF) lacking iron (apo-hTF), which was independently determined by two methods: 1) the crystal structure of recombinant non-glycosylated apo-hTF was solved at 2.7-A resolution using a multiple wavelength anomalous dispersion phasing strategy, by substituting the nine methionines in hTF with selenomethionine and 2) the structure of glycosylated apo-hTF (isolated from serum) was determined to a resolution of 2.7A by molecular replacement using the human apo-N-lobe and the rabbit holo-C1-subdomain as search models. These two crystal structures are essentially identical. They represent the first published model for full-length human transferrin and reveal that, in contrast to family members (human lactoferrin and hen ovotransferrin), both lobes are almost equally open: 59.4 degrees and 49.5 degrees rotations are required to open the N- and C-lobes, respectively (compared with closed pig TF). Availability of this structure is critical to a complete understanding of the metal binding properties of each lobe of hTF; the apo-hTF structure suggests that differences in the hinge regions of the N- and C-lobes may influence the rates of iron binding and release. In addition, we evaluate potential interactions between apo-hTF and the human transferrin receptor.
- NIDDK, National Institutes of Health, Bethesda, Maryland 20892, USA.
Organizational Affiliation: