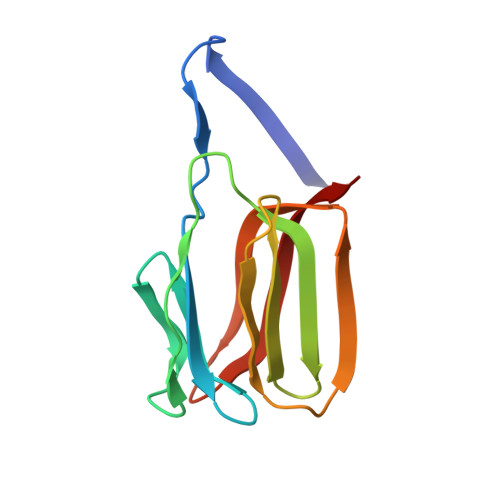
Domain-swapped structure of the potent antiviral protein griffithsin and its mode of carbohydrate binding.
Ziolkowska, N.E., O'keefe, B.R., Mori, T., Zhu, C., Giomarelli, B., Vojdani, F., Palmer, K.E., McMahon, J.B., Wlodawer, A.(2006) Structure 14: 1127-1135
- PubMed: 16843894 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2006.05.017
- Primary Citation Related Structures:
2GTY, 2GUC, 2GUD, 2GUE, 2GUX - PubMed Abstract:
The crystal structure of griffithsin, an antiviral lectin from the red alga Griffithsia sp., was solved and refined at 1.3 A resolution for the free protein and 0.94 A for a complex with mannose. Griffithsin molecules form a domain-swapped dimer, in which two beta strands of one molecule complete a beta prism consisting of three four-stranded sheets, with an approximate 3-fold axis, of another molecule. The structure of each monomer bears close resemblance to jacalin-related lectins, but its dimeric structure is unique. The structures of complexes of griffithsin with mannose and N-acetylglucosamine defined the locations of three almost identical carbohydrate binding sites on each monomer. We have also shown that griffithsin is a potent inhibitor of the coronavirus responsible for severe acute respiratory syndrome (SARS). Antiviral potency of griffithsin is likely due to the presence of multiple, similar sugar binding sites that provide redundant attachment points for complex carbohydrate molecules present on viral envelopes.
- Protein Structure Section, Macromolecular Crystallography Laboratory, National Cancer Institute, NCI-Frederick, Frederick, MD 21702, USA.
Organizational Affiliation: