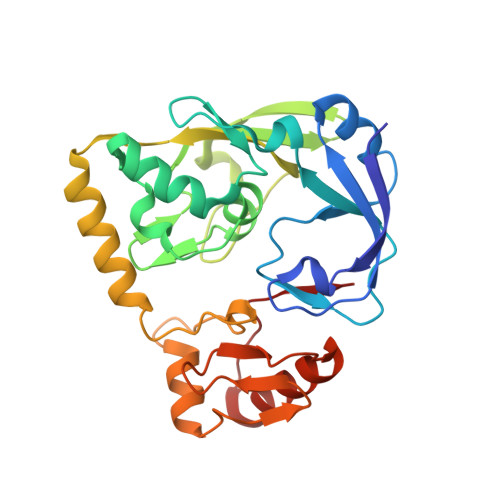
Crystal structure of UDP-N-acetylenolpyruvylglucosamine reductase (MurB) from Thermus caldophilus
Kim, M.-K., Cho, M.K., Song, H.-E., Kim, D., Park, B.-H., Lee, J.H., Kang, G.B., Kim, S.H., Im, Y.J., Lee, D.-S., Eom, S.H.(2006) Proteins 66: 751-754
- PubMed: 17120230 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21174
- Primary Citation Related Structures:
2GQT, 2GQU - Department of Life Science, Gwangju Institute of Science and Technology, Gwangju 500-712, Korea.
Organizational Affiliation: