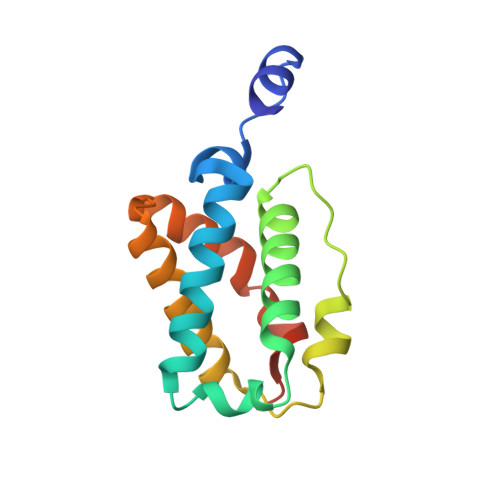
Ligand interactions in the distal heme pocket of Mycobacterium tuberculosis truncated hemoglobin N: roles of TyrB10 and GlnE11 residues
Ouellet, Y., Milani, M., Couture, M., Bolognesi, M., Guertin, M.(2006) Biochemistry 45: 8770-8781
- PubMed: 16846220 Search on PubMed
- DOI: https://doi.org/10.1021/bi060112o
- Primary Citation Related Structures:
2GKM, 2GKN, 2GL3, 2GLN - PubMed Abstract:
The crystallographic structure of oxygenated trHbN from Mycobacterium tuberculosis showed an extended heme distal site hydrogen-bonding network that includes Y(B10), Q(E11), and the bound O(2) (Milani, M., et al. (2001) EMBO J. 20, 3902-3909). In the present work, we analyze the effects that substitutions at the B10 and E11 positions exert on the heme and its coordinated ligands, using steady-state resonance Raman spectroscopy, absorption spectroscopy and X-ray crystallography. Our results show that (1) residues Y(B10) and Q(E11) control the binding and the ionization state of the heme-bound water molecules in ferric trHbN and are important in keeping the sixth coordination position vacant in deoxy trHbN; (2) residue Q(E11) plays a role in maintaining the integrity of the proximal Fe-His bond in deoxy trHbN; (3) in wild-type oxy-trHbN, the size and hydrogen-bonding capability of residue E11 is important to sustain proper interaction between Y(B10) and the heme-bound O(2); (4) CO-trHbN is in a conformational equilibrium, where either the Y(B10) or the Q(E11) residue interacts with the heme-bound CO; and (5) Y(B10) and Q(E11) residues control the conformation (and likely the dynamics) of the protein matrix tunnel gating residue F(E15). These findings suggest that the functional processes of ligand binding and diffusion are controlled in trHbN through the dynamic interaction of residues Y(B10), Q(E11), F(E15), and the heme ligand.
- Department of Biochemistry and Microbiology, Laval University, Quebec, Canada.
Organizational Affiliation: