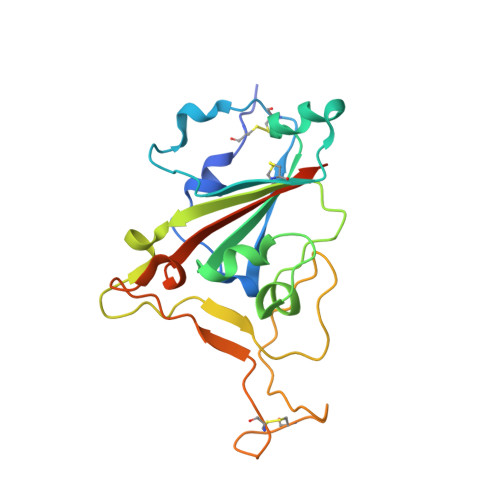
Structural basis of neutralization by a human anti-severe acute respiratory syndrome spike protein antibody, 80R.
Hwang, W.C., Lin, Y., Santelli, E., Sui, J., Jaroszewski, L., Stec, B., Farzan, M., Marasco, W.A., Liddington, R.C.(2006) J Biological Chem 281: 34610-34616
- PubMed: 16954221 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M603275200
- Primary Citation Related Structures:
2GHV, 2GHW - PubMed Abstract:
Severe acute respiratory syndrome (SARS) is a newly emerged infectious disease that caused pandemic spread in 2003. The etiological agent of SARS is a novel coronavirus (SARS-CoV). The coronaviral surface spike protein S is a type I transmembrane glycoprotein that mediates initial host binding via the cell surface receptor angiotensin-converting enzyme 2 (ACE2), as well as the subsequent membrane fusion events required for cell entry. Here we report the crystal structure of the S1 receptor binding domain (RBD) in complex with a neutralizing antibody, 80R, at 2.3 A resolution, as well as the structure of the uncomplexed S1 RBD at 2.2 A resolution. We show that the 80R-binding epitope on the S1 RBD overlaps very closely with the ACE2-binding site, providing a rationale for the strong binding and broad neutralizing ability of the antibody. We provide a structural basis for the differential effects of certain mutations in the spike protein on 80R versus ACE2 binding, including escape mutants, which should facilitate the design of immunotherapeutics to treat a future SARS outbreak. We further show that the RBD of S1 forms dimers via an extensive interface that is disrupted in receptor- and antibody-bound crystal structures, and we propose a role for the dimer in virus stability and infectivity.
- Infectious and Inflammatory Disease Center, Burnham Institute for Medical Research, La Jolla, California 92037, USA.
Organizational Affiliation: