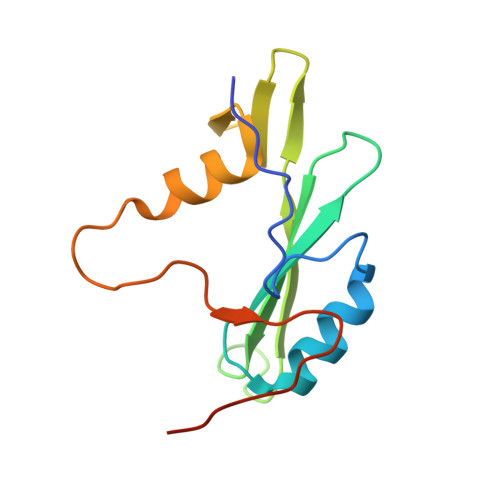
Solution structure and phosphopeptide binding of the SH2 domain from the human Bruton's tyrosine kinase
Huang, K.-C., Cheng, H.-T., Pai, M.-T., Tzeng, S.-R., Cheng, J.-W.(2006) J Biomol NMR 36: 73-78
- PubMed: 16969585 Search on PubMed
- DOI: https://doi.org/10.1007/s10858-006-9064-3
- Primary Citation Related Structures:
2GE9 - Institute of Biotechnology and Department of Life Science, National Tsing Hua University, Hsinchu, Taiwan, ROC.
Organizational Affiliation: