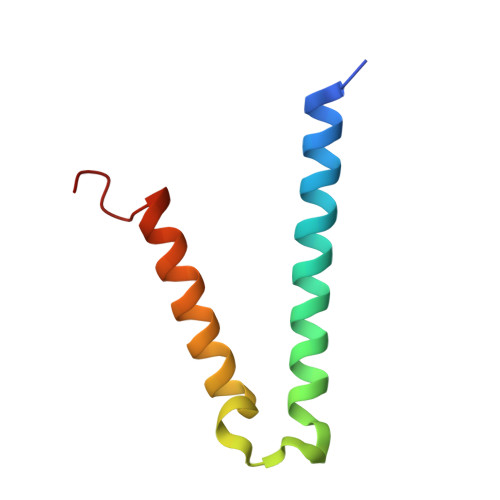
Protonation-mediated structural flexibility in the F conjugation regulatory protein, TraM.
Lu, J., Edwards, R.A., Wong, J.J., Manchak, J., Scott, P.G., Frost, L.S., Glover, J.N.(2006) EMBO J 25: 2930-2939
- PubMed: 16710295 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7601151
- Primary Citation Related Structures:
2G7O, 2G9E - PubMed Abstract:
TraM is essential for F plasmid-mediated bacterial conjugation, where it binds to the plasmid DNA near the origin of transfer, and recognizes a component of the transmembrane DNA transfer complex, TraD. Here we report the 1.40 A crystal structure of the TraM core tetramer (TraM58-127). TraM58-127 is a compact eight-helical bundle, in which the N-terminal helices from each protomer interact to form a central, parallel four-stranded coiled-coil, whereas each C-terminal helix packs in an antiparallel arrangement around the outside of the structure. Four protonated glutamic acid residues (Glu88) are packed in a hydrogen-bonded arrangement within the central four-helix bundle. Mutational and biophysical analyses indicate that this protonated state is in equilibrium with a deprotonated tetrameric form characterized by a lower helical content at physiological pH and temperature. Comparison of TraM to its Glu88 mutants predicted to stabilize the helical structure suggests that the protonated state is the active form for binding TraD in conjugation.
- Department of Biochemistry, University of Alberta, Edmonton, Alberta, Canada.
Organizational Affiliation: