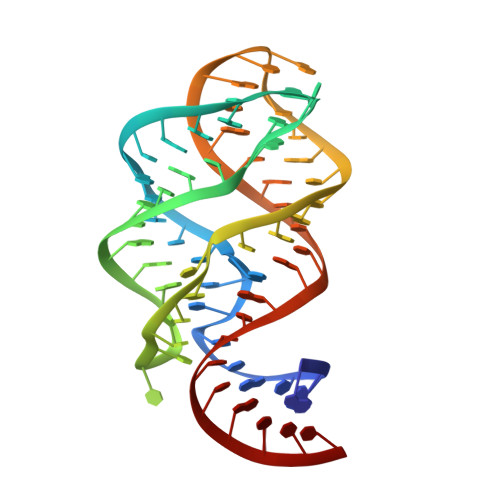
Modified pyrimidines specifically bind the purine riboswitch.
Gilbert, S.D., Mediatore, S.J., Batey, R.T.(2006) J Am Chem Soc 128: 14214-14215
- PubMed: 17076468 Search on PubMed
- DOI: https://doi.org/10.1021/ja063645t
- Primary Citation Related Structures:
2G9C - PubMed Abstract:
The purine riboswitch is a genetic regulatory element found in the 5'-untranslated regions of Gram-positive bacteria that regulates expression of the mRNA specifically in response to either guanine or adenine. We report that the adenine-responsive RNA element is also capable of specifically recognizing pyrimidine compounds bearing modifications at the 6- or 5,6-positions in a fashion similar to that of purine compounds. Using isothermal titration calorimetry and X-ray crystallography, the binding of these compounds is characterized.
- Department of Chemistry and Biochemistry, University of Colorado at Boulder, Campus Box 215, Boulder, Colorado 80309-0215, USA.
Organizational Affiliation: