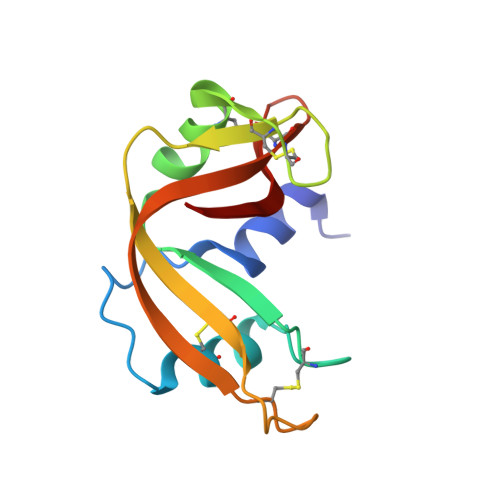
The binding of 3'-N-piperidine-4-carboxyl-3'-deoxy-ara-uridine to ribonuclease A in the crystal.
Leonidas, D.D., Maiti, T.K., Samanta, A., Dasgupta, S., Pathak, T., Zographos, S.E., Oikonomakos, N.G.(2006) Bioorg Med Chem 14: 6055-6064
- PubMed: 16730994 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmc.2006.05.011
- Primary Citation Related Structures:
2G8Q, 2G8R - PubMed Abstract:
The binding of a moderate inhibitor, 3'-N-piperidine-4-carboxyl-3'-deoxy-ara-uridine, to ribonuclease A has been studied by X-ray crystallography at 1.7A resolution. Two inhibitor molecules are bound in the central RNA binding cavity of RNase A exploiting interactions with residues from peripheral binding sites rather than from the active site of the enzyme. The uracyl moiety of the first inhibitor molecule occupies the purine-preferring site of RNase A, while the rest of the molecule projects to the solvent. The second inhibitor molecule binds with the carboxyl group at the pyrimidine recognition site and the uridine moiety exploits interactions with RNase A residues Lys66, His119 and Asp121. Comparative structural analysis of the 3'-N-piperidine-4-carboxyl-3'-deoxy-ara-uridine complex with other RNase A-ligand complexes provides a structural explanation of its potency. The crystal structure of the RNase A-3'-N-piperidine-4-carboxyl-3'-deoxy-ara-uridine complex provides evidence of a novel ligand-binding pattern in RNase A for 3'-N-aminonucleosides that was not anticipated by modelling studies, while it also suggests ways to improve the efficiency and selectivity of such compounds to develop pharmaceuticals against pathologies associated with RNase A homologues.
- Institute of Organic and Pharmaceutical Chemistry, The National Hellenic Research Foundation, 48 Vas. Constantinou Avenue, 11635 Athens, Greece. ddl@eie.gr
Organizational Affiliation: