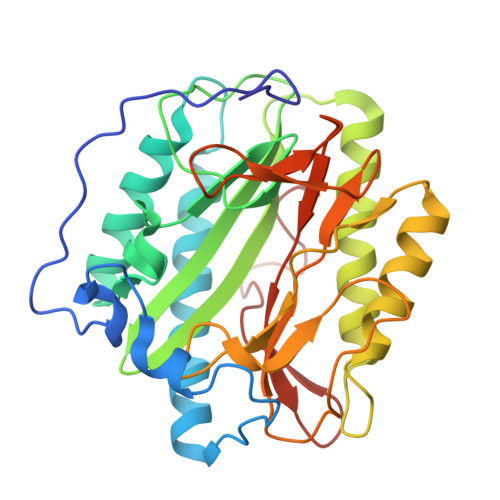
Identification of Pyridinylpyrimidines as Inhibitors of Human Methionine Aminopeptidases.
Hu, X., Addlagatta, A., Matthews, B.W., Liu, J.O.(2006) Angew Chem Int Ed Engl 45: 3772-3775
- PubMed: 16724298 Search on PubMed
- DOI: https://doi.org/10.1002/anie.200600757
- Primary Citation Related Structures:
2G6P - Department of Pharmacology and Molecular Sciences, Johns Hopkins University School of Medicine, 725 N. Wolfe Street, Baltimore, MD 21205, USA.
Organizational Affiliation: