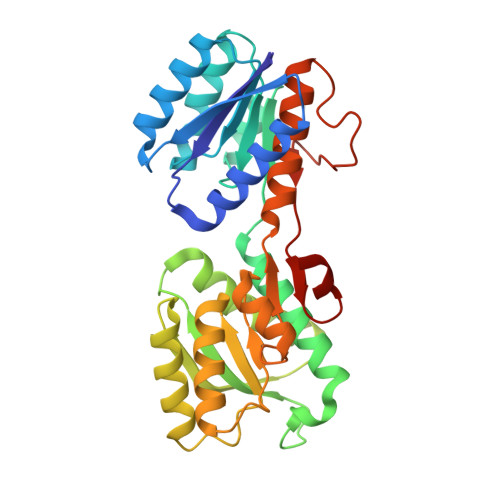
Conformational changes of glucose/galactose-binding protein illuminated by open, unliganded, and ultra-high-resolution ligand-bound structures.
Borrok, M.J., Kiessling, L.L., Forest, K.T.(2007) Protein Sci 16: 1032-1041
- PubMed: 17473016 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.062707807
- Primary Citation Related Structures:
2FVY, 2FW0 - PubMed Abstract:
D-Glucose/D-Galactose-binding protein (GGBP) mediates chemotaxis toward and active transport of glucose and galactose in a number of bacterial species. GGBP, like other periplasmic binding proteins, can exist in open (ligand-free) and closed (ligand-bound) states. We report a 0.92 angstroms resolution structure of GGBP from Escherichia coli in the glucose-bound state and the first structure of an open, unbound form of GGBP (at 1.55 angstroms resolution). These structures vary in the angle between the two structural domains; the observed difference of 31 degrees arises from torsion angle changes in a three-segment hinge. A comparison with the closely related periplasmic receptors, ribose- and allose-binding proteins, shows that the GGBP hinge residue positions that undergo the largest conformational changes are different. Furthermore, the high-quality data collected for the atomic resolution glucose-bound structure allow for the refinement of specific hydrogen atom positions, the assignment of alternate side chain conformations, the first description of CO(2) trapped after radiation-induced decarboxylation, and insight into the role of the exo-anomeric effect in sugar binding. Together, these structures provide insight into how the hinge-bending movement of GGBP facilitates ligand binding, transport, and signaling.
- Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53706, USA.
Organizational Affiliation: