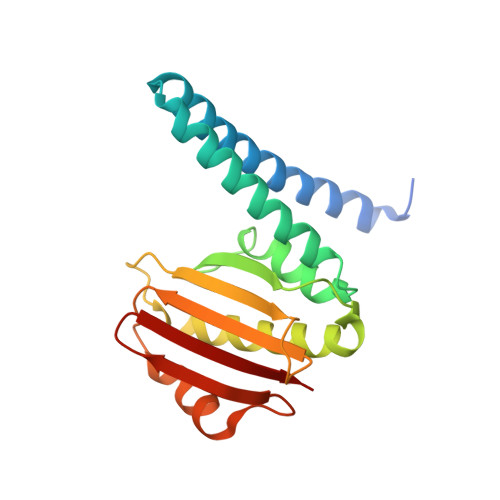
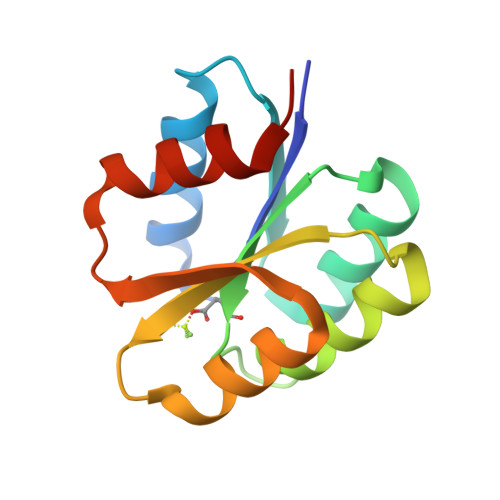
The Crystal Structure of Beryllofluoride Spo0F in Complex with the Phosphotransferase Spo0B Represents a Phosphotransfer Pretransition State.
Varughese, K.I., Tsigelny, I., Zhao, H.(2006) J Bacteriol 188: 4970-4977
- PubMed: 16788205 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.00160-06
- Primary Citation Related Structures:
2FTK - PubMed Abstract:
A number of regulatory circuits in biological systems function through the exchange of phosphoryl groups from one protein to another. Spo0F and Spo0B are components of a phosphorelay that control sporulation in the bacterium Bacillus subtilis through the exchange of a phosphoryl group. Using beryllofluoride as a mimic for phosphorylation, we trapped the interaction of the phosphorylated Spo0F with Spo0B in the crystal lattice. The transition state of phosphoryl transfer continues to be a highly debated issue, as to whether it is associative or dissociative in nature. The geometry of Spo0F binding to Spo0B favors an associative mechanism for phosphoryl transfer. In order to visualize the autophosphorylation of the histidine kinase, KinA, and the subsequent phosphoryl transfer to Spo0F, we generated in silico models representing these reaction steps.
- Division of Cellular Biology, Department of Molecular and Experimental Medicine, MEM-116, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA. kiv@scripps.edu
Organizational Affiliation: