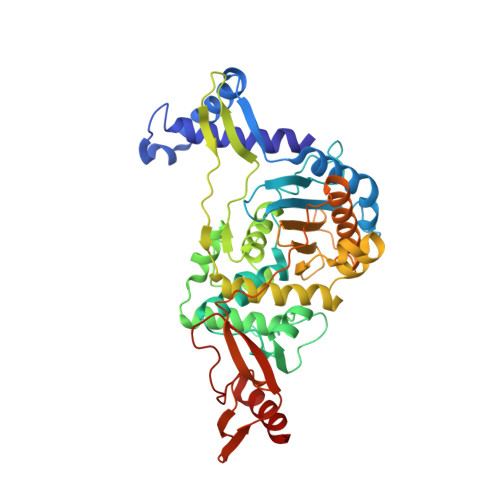
Structural analysis of an "open" form of PBP1B from Streptococcus pneumoniae.
Lovering, A.L., De Castro, L., Lim, D., Strynadka, N.C.(2006) Protein Sci 15: 1701-1709
- PubMed: 16751607 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.062112106
- Primary Citation Related Structures:
2FFF - PubMed Abstract:
The class A PBP1b from Streptococcus pneumoniae is responsible for glycosyltransferase and transpeptidase (TP) reactions, forming the peptidoglycan of the bacterial cell wall. The enzyme has been produced in a stable, soluble form and undergoes time-dependent proteolysis to leave an intact TP domain. Crystals of this TP domain were obtained, diffracting to 2.2 A resolution, and the structure was solved by using molecular replacement. Analysis of the structure revealed an "open" active site, with important conformational differences to the previously determined "closed" apoenzyme. The active-site nucleophile, Ser460, is in an orientation that allows for acylation by beta-lactams. Consistent with the productive conformation of the conserved active-site catalytic residues, adjacent loops show only minor deviation from those of known acyl-enzyme structures. These findings are discussed in the context of enzyme functionality and the possible conformational sampling of PBP1b between active and inactive states.
- Department of Biochemistry and Molecular Biology, Center for Blood Research, University of British Columbia, Vancouver, Canada.
Organizational Affiliation: