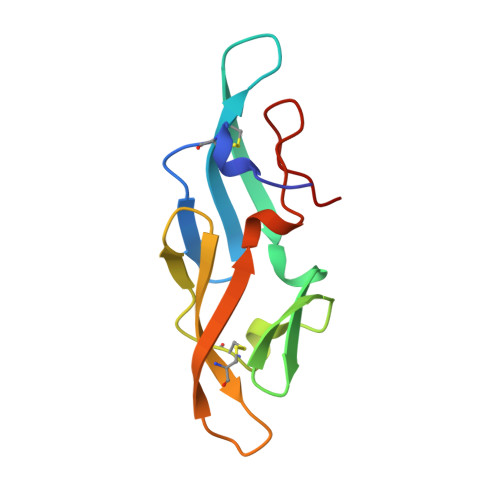
Solution structure of cyanovirin-N, a potent HIV-inactivating protein.
Bewley, C.A., Gustafson, K.R., Boyd, M.R., Covell, D.G., Bax, A., Clore, G.M., Gronenborn, A.M.(1998) Nat Struct Biol 5: 571-578
- PubMed: 9665171 Search on PubMed
- DOI: https://doi.org/10.1038/828
- Primary Citation Related Structures:
2EZM, 2EZN - PubMed Abstract:
The solution structure of cyanovirin-N, a potent 11,000 Mr HIV-inactivating protein that binds with high affinity and specificity to the HIV surface envelope protein gp120, has been solved by nuclear magnetic resonance spectroscopy, including extensive use of dipolar couplings which provide a priori long range structural information. Cyanovirin-N is an elongated, largely beta-sheet protein that displays internal two-fold pseudosymmetry. The two sequence repeats (residues 1-50 and 51-101) share 32% sequence identity and superimpose with a backbone atomic root-mean-square difference of 1.3 A. The two repeats, however, do not form separate domains since the overall fold is dependent on numerous contacts between them. Rather, two symmetrically related domains are formed by strand exchange between the two repeats. Analysis of surface hydrophobic clusters suggests the location of potential binding sites for protein-protein interactions.
- Laboratory of Chemical Physics, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, Maryland 20892-0520, USA.
Organizational Affiliation: