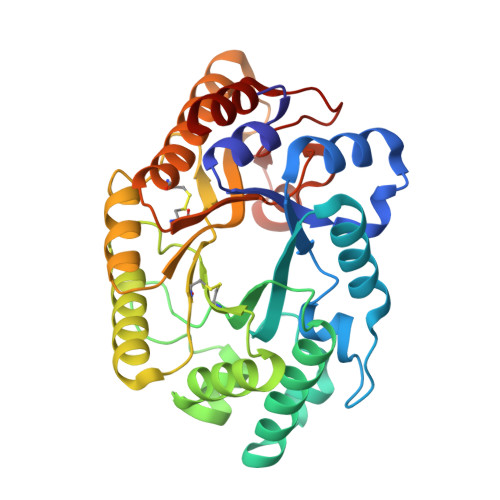
Crystal structure of the catalytic domain of the beta-1,4-glycanase cex from Cellulomonas fimi.
White, A., Withers, S.G., Gilkes, N.R., Rose, D.R.(1994) Biochemistry 33: 12546-12552
- PubMed: 7918478 Search on PubMed
- DOI: https://doi.org/10.1021/bi00208a003
- Primary Citation Related Structures:
2EXO - PubMed Abstract:
beta-1,4-Glycanases, principally cellulases and xylanases, are responsible for the hydrolysis of plant biomass. The bifunctional beta-1,4-xylanase/glucanase Cex from the bacterium Cellulomonas fimi, one of a large family of cellulases/xylanases, depolymerizes oligosaccharides and releases a disaccharide unit from the substrate nonreducing end. Hydrolysis occurs with net retention of the anomeric configuration of the sugar through a double-displacement mechanism involving a covalent glycosyl-enzyme intermediate. The active site nucleophile, Glu233, has been unambiguously identified by trapping of such an intermediate [Tull et al. (1991) J. Biol. Chem. 266, 15621-15625] and the acid/base catalyst, Glu127, by detailed kinetic analysis of mutants [MacLeod et al. (1994) Biochemistry 33, 6371-6376]. However, little is known about the enzyme's overall folding and its active site architecture. We report here the high-resolution crystal structure of the catalytic domain of Cex. The atomic structure refinement results in a model that includes 2400 protein atoms and 45 water molecules, with an R-factor of 0.217 for data extending to 1.8-A resolution. The protein forms an eight-parallel-stranded alpha/beta-barrel, which is a novel folding pattern for a microbial beta-glycanase. The active site, inferred from the location of Glu233, Glu127, and other conserved residues, is an open cleft on the carboxy-terminal end of the alpha/beta-barrel. An extensive hydrogen-bonding network stabilizes the ionization states of the key residues; in particular, the Asp235-His205-Glu233 hydrogen-bonding network may play a role in modulating the ionization state of Glu233 and in controlling local charge balance during the reaction.
- Protein Engineering Network of Centres of Excellence, Ontario Cancer Institute, University of Toronto, Canada.
Organizational Affiliation: