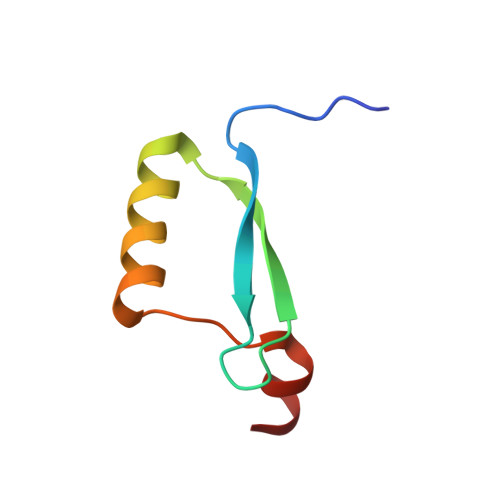
Solution structure of the partially folded high-risk human papilloma virus 45 oncoprotein E7.
Ohlenschlager, O., Seiboth, T., Zengerling, H., Briese, L., Marchanka, A., Ramachandran, R., Baum, M., Korbas, M., Meyer-Klaucke, W., Durst, M., Gorlach, M.(2006) Oncogene 25: 5953-5959
- PubMed: 16636661 Search on PubMed
- DOI: https://doi.org/10.1038/sj.onc.1209584
- Primary Citation Related Structures:
2EWL, 2F8B - PubMed Abstract:
The oncoprotein E7 of human papilloma viruses (HPV) is involved in the pathogenesis and maintenance of human cervical cancers. The most prevalent HPV types found in cervix carcinomas are HPV16, 18 and 45. The structure of the E7 dimer from HPV45 (PDB 2F8B) was determined by nuclear magnetic resonance spectroscopy. Each monomer comprises an unfolded N-terminus and a well-structured C-terminal domain with a beta1beta2alpha1beta3alpha2 topology representing a unique zinc-binding fold found only for E7. Dimerization occurs through the alpha1/alpha1' helices and intermolecular beta-sheet formation but excludes the zinc-binding sites. E7 is reported to interact with a number of cellular proteins (e.g. pRb, p21(CIP1)). Binding of a peptide derived from the C-terminus of p21(CIP1) to the C-terminal domain of E7 was characterized by monitoring chemical shift perturbations of the amide groups of E7. This provides direct evidence that a shallow groove situated between alpha1 and beta1 of the E7 C-terminal domain is interacting with the C-terminus of p21(CIP1). Intriguingly, this binding site overlaps with the low-affinity binding site on E7 for the C-domain of pRb.
- Fritz-Lipmann-Institut, Jena, Germany.
Organizational Affiliation: