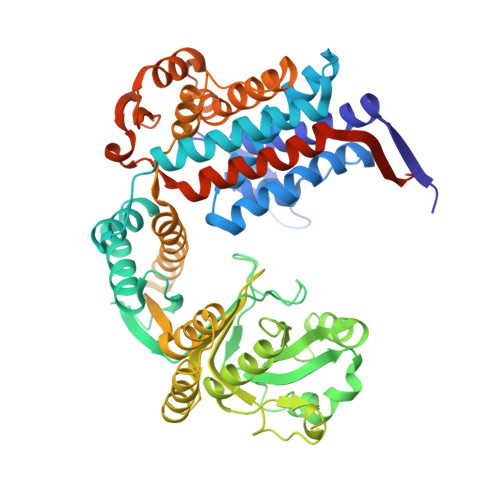
Crystal structure of the temperature-sensitive and allosteric-defective chaperonin GroEL(E461K).
Cabo-Bilbao, A., Spinelli, S., Sot, B., Agirre, J., Mechaly, A.E., Muga, A., Guerin, D.M.A.(2006) J Struct Biol 155: 482-492
- PubMed: 16904907 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2006.06.008
- Primary Citation Related Structures:
2EU1, 2YEY - PubMed Abstract:
The chaperonin GroEL adopts a double-ring structure with various modes of allosteric communication. The simultaneous positive intra-ring and negative inter-ring co-operativities alternate the functionality of the folding cavities in both protein rings. Negative inter-ring co-operativity is maintained through different inter-ring interactions, including a salt bridge involving Glu 461. Replacement of this residue by Lys modifies the temperature sensitivity of the substrate-folding activity of this protein, most likely as a result of the loss of inter-ring co-operativity. The crystal structure of the mutant chaperonin GroELE461K has been determined at 3.3A and compared with other structures: the wild-type GroEL, an allosteric defective GroEL double mutant and the GroEL-GroES-(ADP)7 complex. The inter-ring region of the mutant exhibits the following characteristics: (i) no salt-bridge stabilizes the inter-ring interface; (ii) the mutated residue plays a central role in defining the relative ring rotation (of about 22 degrees) around the 7-fold axis; (iii) an increase in the inter-ring distance and solvent accessibility of the inter-ring interface; and (iv) a 2-fold reduction in the stabilization energy of the inter-ring interface, due to the modification of inter-ring interactions. These characteristics explain how the thermal sensitivity of the protein's fundamental properties permits GroEL to distinguish physiological (37 degrees C) from stress (42 degrees C) temperatures.
- Unidad de Biofísica (CSIC-UPV/EHU), P.O. Box 644, E-48080 Bilbao, Spain.
Organizational Affiliation: