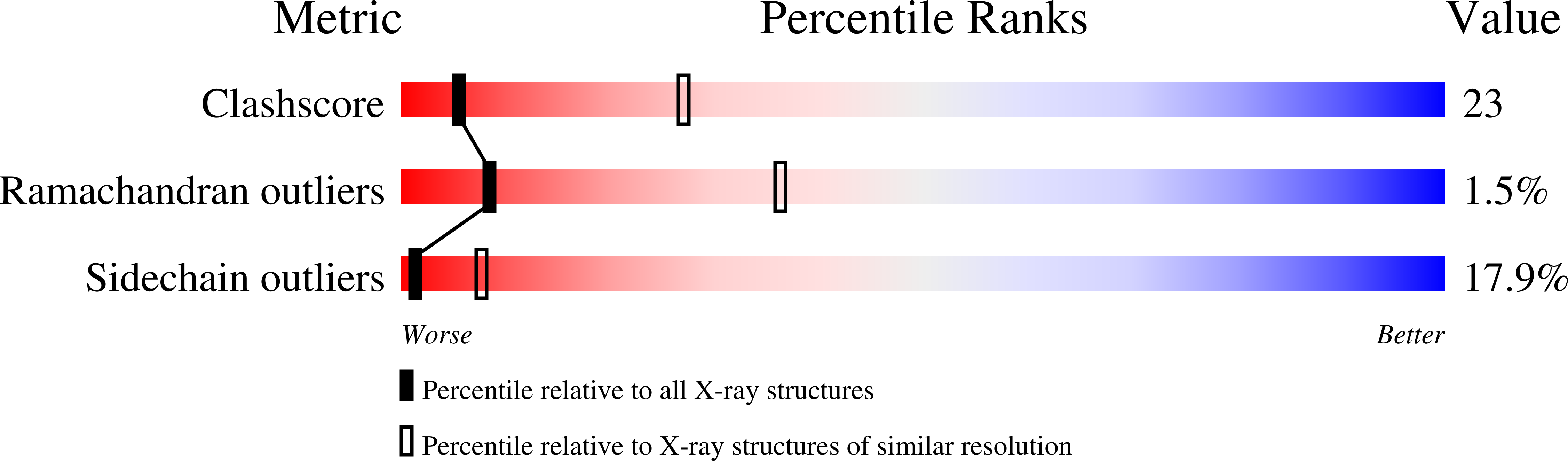X-ray studies of aspartic proteinase-statine inhibitor complexes.
Cooper, J.B., Foundling, S.I., Blundell, T.L., Boger, J., Jupp, R.A., Kay, J.(1989) Biochemistry 28: 8596-8603
- PubMed: 2690945 Search on PubMed
- DOI: https://doi.org/10.1021/bi00447a049
- Primary Citation Related Structures:
2ER0, 2ER9 - PubMed Abstract:
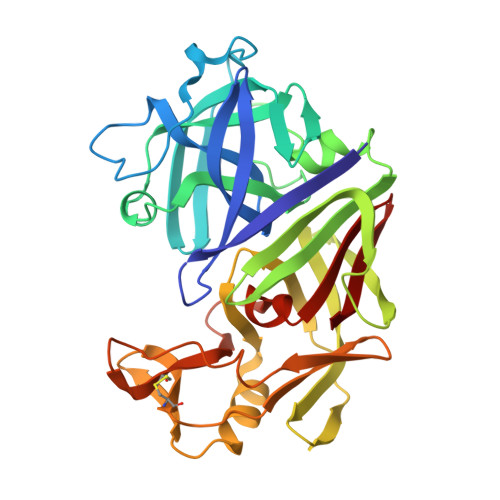
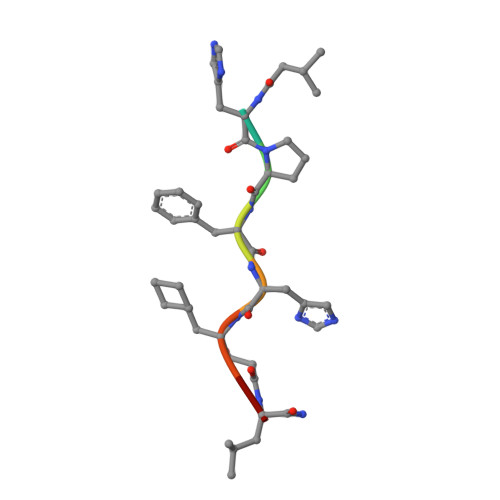
The conformation of a statine-containing renin inhibitor complexed with the aspartic proteinase from the fungus Endothia parasitica (EC 3.4.23.6) has been determined by X-ray diffraction at 2.2-A resolution (R = 0.17). We describe the structure of the complex at high resolution and compare this with a 3.0-A resolution analysis of a bound inhibitor, L-364,099, containing a cyclohexylalanine analogue of statine. The inhibitors bind in extended conformations in the long active-site cleft, and the hydroxyl of the transition-state analogue, statine, interacts strongly with the catalytic aspartates via hydrogen bonds to the essential carboxyl groups. This work provides a detailed structural analysis of the role of statine in peptide inhibitors. It shows conclusively that statine should be considered a dipeptide analogue (occupying P1 to P1') despite lacking the equivalent of a P1' side chain, although other inhibitor residues (especially P2) may compensate by interacting at the unoccupied S1' specificity subsite.
- Department of Crystallography, Birkbeck College, London, U.K.
Organizational Affiliation: