Structural characterization of a novel bacterial aminopeptidase with an apical domain from aneurinibacillus sp. strain AM-1
Akioka, M., Nakano, H., Tsujimoto, Y., Matsui, H., Nakatsu, T., Kato, H., Watanabe, K.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Aminopeptidase | 421 | Aneurinibacillus sp. AM-1 | Mutation(s): 0 | 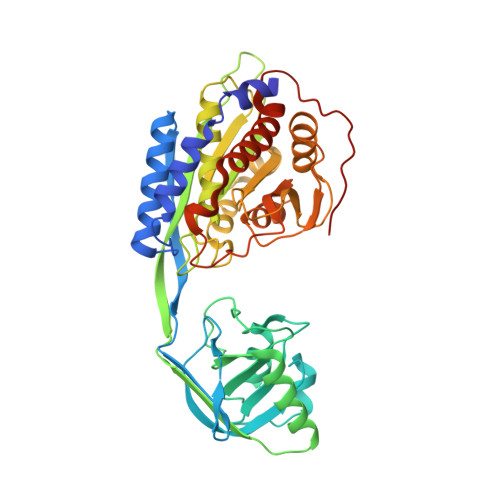 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A2V759 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| BES Download:Ideal Coordinates CCD File | H [auth A] | 2-(3-AMINO-2-HYDROXY-4-PHENYL-BUTYRYLAMINO)-4-METHYL-PENTANOIC ACID C16 H24 N2 O4 VGGGPCQERPFHOB-RDBSUJKOSA-N |  | ||
| ZN Download:Ideal Coordinates CCD File | B [auth A] C [auth A] D [auth A] E [auth A] F [auth A] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| IPA Download:Ideal Coordinates CCD File | I [auth A] | ISOPROPYL ALCOHOL C3 H8 O KFZMGEQAYNKOFK-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 93.813 | α = 90 |
| b = 68.678 | β = 90 |
| c = 77.044 | γ = 90 |
| Software Name | Purpose |
|---|---|
| CNS | refinement |
| CrystalClear | data collection |
| MOSFLM | data reduction |
| SCALA | data scaling |