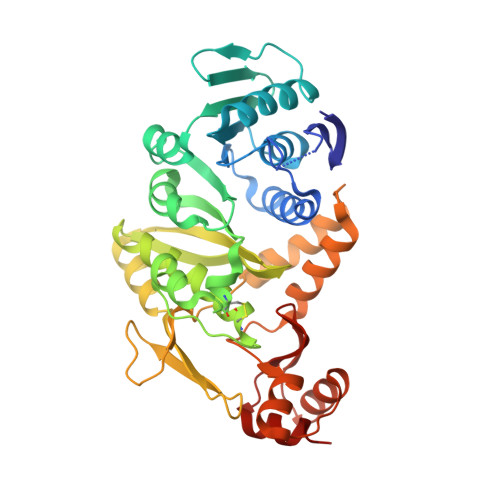
Crystal Structure of tRNA N(2),N(2)-Guanosine Dimethyltransferase Trm1 from Pyrococcus horikoshii
Ihsanawati, Nishimoto, M., Higashijima, K., Shirouzu, M., Grosjean, H., Bessho, Y., Yokoyama, S.(2008) J Mol Biology 383: 871-884
- PubMed: 18789948 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.08.068
- Primary Citation Related Structures:
2DUL, 2EJT, 2EJU, 2YTZ - PubMed Abstract:
Trm1 catalyzes a two-step reaction, leading to mono- and dimethylation of guanosine at position 26 in most eukaryotic and archaeal tRNAs. We report the crystal structures of Trm1 from Pyrococcus horikoshii liganded with S-adenosyl-l-methionine or S-adenosyl-l-homocysteine. The protein comprises N-terminal and C-terminal domains with class I methyltransferase and novel folds, respectively. The methyl moiety of S-adenosyl-l-methionine points toward the invariant Phe27 and Phe140 within a narrow pocket, where the target G26 might flip in. Mutagenesis of Phe27 or Phe140 to alanine abolished the enzyme activity, indicating their role in methylating G26. Structural analyses revealed that the movements of Phe140 and the loop preceding Phe27 may be involved in dissociation of the monomethylated tRNA*Trm1 complex prior to the second methylation. Moreover, the catalytic residues Asp138, Pro139, and Phe140 are in a different motif from that in DNA 6-methyladenosine methyltransferases, suggesting a different methyl transfer mechanism in the Trm1 family.
- Systems and Structural Biology Center, Yokohama Institute, RIKEN, 1-7-22 Suehiro, Tsurumi, Yokohama 230-0045, Japan.
Organizational Affiliation: