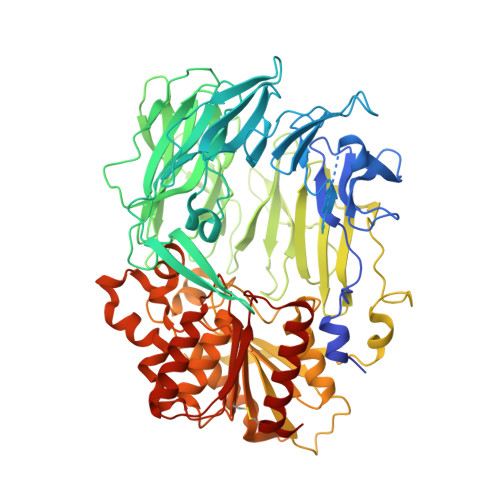
Dipeptidyl aminopeptidase IV from Stenotrophomonas maltophilia exhibits activity against a substrate containing a 4-hydroxyproline residue
Nakajima, Y., Ito, K., Toshima, T., Egawa, T., Zheng, H., Oyama, H., Wu, Y.-F., Takahashi, E., Kyono, K., Yoshimoto, T.(2008) J Bacteriol 190: 7819-7829
- PubMed: 18820015 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.02010-07
- Primary Citation Related Structures:
2ECF - PubMed Abstract:
The crystal structure of dipeptidyl aminopeptidase IV from Stenotrophomonas maltophilia was determined at 2.8-A resolution by the multiple isomorphous replacement method, using platinum and selenomethionine derivatives. The crystals belong to space group P4(3)2(1)2, with unit cell parameters a = b = 105.9 A and c = 161.9 A. Dipeptidyl aminopeptidase IV is a homodimer, and the subunit structure is composed of two domains, namely, N-terminal beta-propeller and C-terminal catalytic domains. At the active site, a hydrophobic pocket to accommodate a proline residue of the substrate is conserved as well as those of mammalian enzymes. Stenotrophomonas dipeptidyl aminopeptidase IV exhibited activity toward a substrate containing a 4-hydroxyproline residue at the second position from the N terminus. In the Stenotrophomonas enzyme, one of the residues composing the hydrophobic pocket at the active site is changed to Asn611 from the corresponding residue of Tyr631 in the porcine enzyme, which showed very low activity against the substrate containing 4-hydroxyproline. The N611Y mutant enzyme was generated by site-directed mutagenesis. The activity of this mutant enzyme toward a substrate containing 4-hydroxyproline decreased to 30.6% of that of the wild-type enzyme. Accordingly, it was considered that Asn611 would be one of the major factors involved in the recognition of substrates containing 4-hydroxyproline.
- Graduate School of Biomedical Sciences, Nagasaki University, 1-14 Bunkyo-machi, Nagasaki 852-8521, Japan.
Organizational Affiliation: