Structural Basis for Flexible Base Recognition by C/Ebpbeta
Tahirov, T.H., Inoue-Bungo, T., Sato, K., Shiina, M., Hamada, K., Ogata, K.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| CCAAT/enhancer-binding protein beta | C [auth A], D [auth B] | 78 | Homo sapiens | Mutation(s): 1 Gene Names: CEBPB, TCF5 |  |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P17676 GTEx: ENSG00000172216 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P17676 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 1 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (5'-D(P*DTP*DAP*DGP*DGP*DAP*DTP*DTP*DGP*DCP*DGP*DCP*DAP*DAP*DTP*DAP*DT)-3') | A [auth C] | 16 | N/A |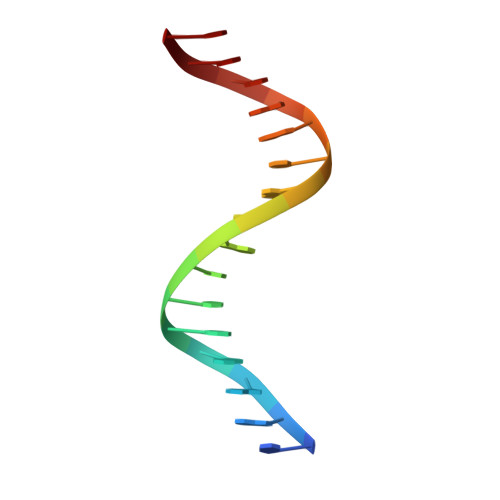  |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
Entity ID: 2 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (5'-D(P*DAP*DAP*DTP*DAP*DTP*DTP*DGP*DCP*DGP*DCP*DAP*DAP*DTP*DCP*DCP*DT)-3') | B [auth D] | 16 | N/A |  |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 101.881 | α = 90 |
| b = 113.319 | β = 90 |
| c = 74.439 | γ = 90 |
| Software Name | Purpose |
|---|---|
| CNS | refinement |
| HKL-2000 | data reduction |
| SCALEPACK | data scaling |
| CNS | phasing |