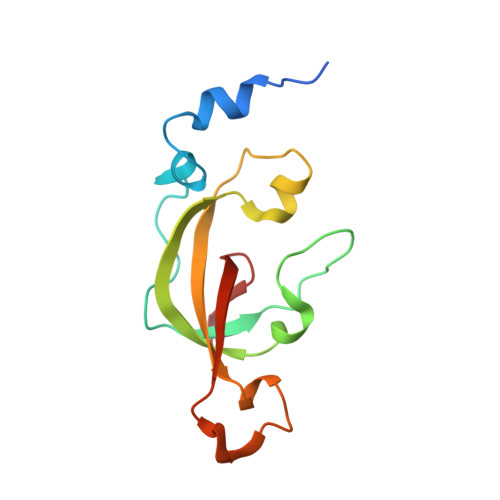
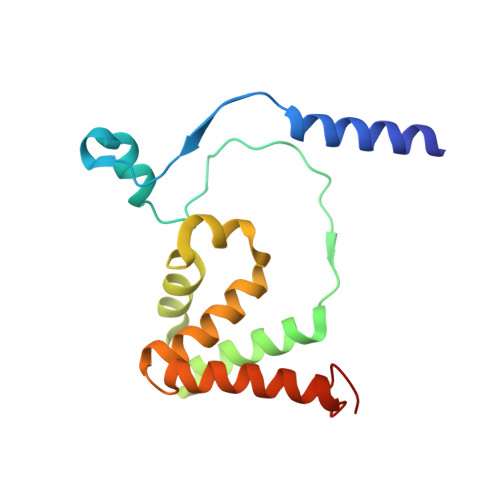
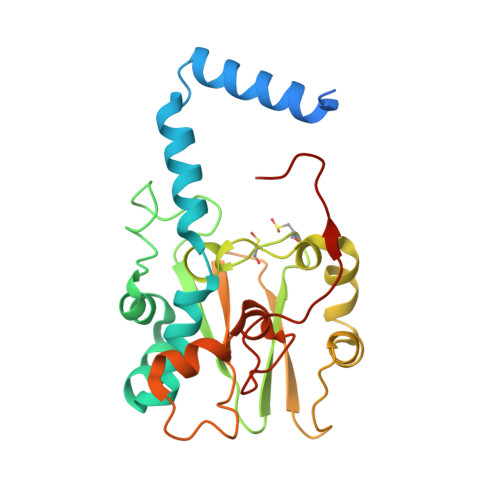
Structural Basis for Catalytic Activation of Thiocyanate Hydrolase Involving Metal-Ligated Cysteine Modification
Arakawa, T., Kawano, Y., Katayama, Y., Nakayama, H., Dohmae, N., Yohda, M., Odaka, M.(2009) J Am Chem Soc 131: 14838-14843
- PubMed: 19785438 Search on PubMed
- DOI: https://doi.org/10.1021/ja903979s
- Primary Citation Related Structures:
2DXB, 2DXC, 2ZZD - PubMed Abstract:
Thiocyanate hydrolase (SCNase) is a member of a family of nitrile hydratase proteins, each of which contains a unique noncorrin cobalt center with two post-translationally modified cysteine ligands, cysteine-sulfenic acid or -sulfenate (Cys-SO(H)), and cysteine-sulfininate (Cys-SO(2)(-)), respectively. We have found that a partially matured recombinant SCNase was activated during storage. The crystal structures of SCNase before and after storage demonstrated that Cys-SO(2)(-) modification of gammaCys131 proceeded to completion prior to storage, while Cys-SO(H) modification of gammaCys133 occurred during storage. SCNase activity was suppressed when gammaCys133 was further oxidized to Cys-SO(2)(-). The correlation between the catalytic activity and the extent of the gammaCys133 modification indicates that the cysteine sulfenic acid modification of gammaCys133 is of primary importance in determining the activity of SCNase.
- Department of Biotechnology and Life Science, Graduate School of Technology, Tokyo University of Agriculture and Technology, Koganei, Tokyo 184-8588, Japan.
Organizational Affiliation: