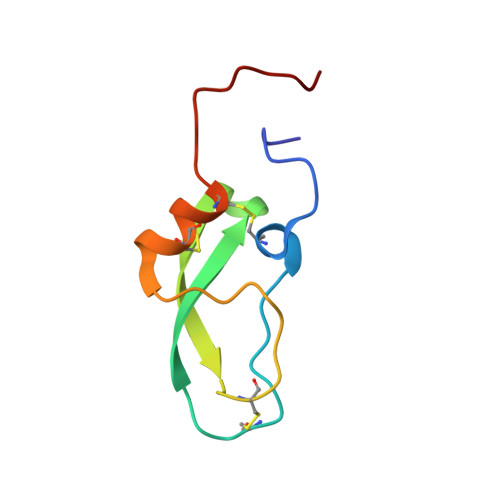
Second Kunitz-type protease inhibitor domain of the human WFIKKN1 protein
Liepinsh, E., Nagy, A., Trexler, M., Patthy, L., Otting, G.(2006) J Biomol NMR 35: 73-78
- PubMed: 16791741 Search on PubMed
- DOI: https://doi.org/10.1007/s10858-006-9013-1
- Primary Citation Related Structures:
2DDI, 2DDJ - Department of Medical Biochemistry and Biophysics, Karolinska Institute, S-17177, Stockholm, Sweden.
Organizational Affiliation: