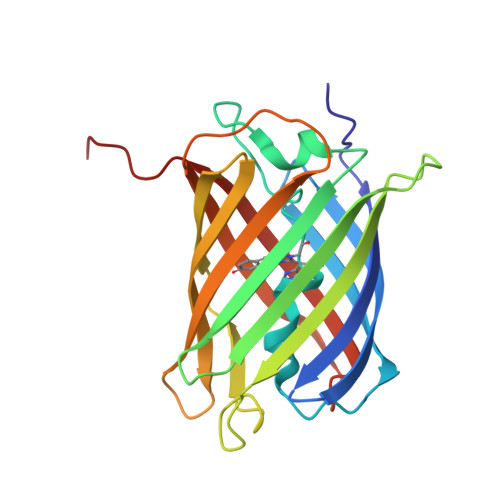
The E1 mechanism in photo-induced beta-elimination reactions for green-to-red conversion of fluorescent proteins.
Tsutsui, H., Shimizu, H., Mizuno, H., Nukina, N., Furuta, T., Miyawaki, A.(2009) Chem Biol 16: 1140-1147
- PubMed: 19942137 Search on PubMed
- DOI: https://doi.org/10.1016/j.chembiol.2009.10.010
- Primary Citation Related Structures:
2DDC, 2DDD - PubMed Abstract:
KikGR is a fluorescent protein engineered to display green-to-red photoconvertibility that is induced by irradiation with ultraviolet or violet light. Similar to Kaede and EosFP, two naturally occurring photoconvertible proteins, KikGR contains a His(62)-Tyr(63)-Gly(64) tripeptide sequence, which forms a green chromophore that can be photoconverted to a red one via formal beta-elimination and subsequent extension of a pi-conjugated system. Using a crystallizable variant of KikGR, we determined the structures of both the green and red state at 1.55 A resolution. The double bond between His(62)-C(alpha) and His(62)-C(beta) in the red chromophore is in a cis configuration, indicating that rotation along the His(62) C(alpha)-C(beta) bond occurs following cleavage of the His(62) N(alpha)-C(alpha) bond. This structural rearrangement provides evidence that the beta-elimination reaction governing the green-to-red photoconversion of KikGR follows an E1 (elimination, unimolecular) mechanism.
- Brain Science Institute, RIKEN, Hirosawa, Wako-city, Saitama, Japan.
Organizational Affiliation: