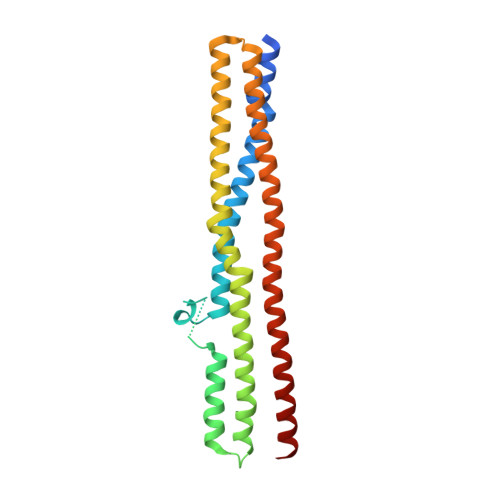
Endophilin BAR domain drives membrane curvature by two newly identified structure-based mechanisms
Masuda, M., Takeda, S., Sone, M., Ohki, T., Mori, H., Kamioka, Y., Mochizuki, N.(2006) EMBO J 25: 2889-2897
- PubMed: 16763557 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7601176
- Primary Citation Related Structures:
1X03, 1X04, 2D4C - PubMed Abstract:
The crescent-shaped BAR (Bin/Amphiphysin/Rvs-homology) domain dimer is a versatile protein module that senses and generates positive membrane curvature. The BAR domain dimer of human endophilin-A1, solved at 3.1 A, has a unique structure consisting of a pair of helix-loop appendages sprouting out from the crescent. The appendage's short helices form a hydrophobic ridge, which runs across the concave surface at its center. Examining liposome binding and tubulation in vitro using purified BAR domain and its mutants indicated that the ridge penetrates into the membrane bilayer and enhances liposome tubulation. BAR domain-expressing cells exhibited marked plasma membrane tubulation in vivo. Furthermore, a swinging-arm mutant lost liposome tubulation activity yet retaining liposome binding. These data suggested that the rigid crescent dimer shape is crucial for the tubulation. We here propose that the BAR domain drives membrane curvature by coordinate action of the crescent's scaffold mechanism and the ridge's membrane insertion in addition to membrane binding via amino-terminal amphipathic helix.
- Department of Structural Analysis, National Cardiovascular Center Research Institute, Suita, Osaka, Japan.
Organizational Affiliation: