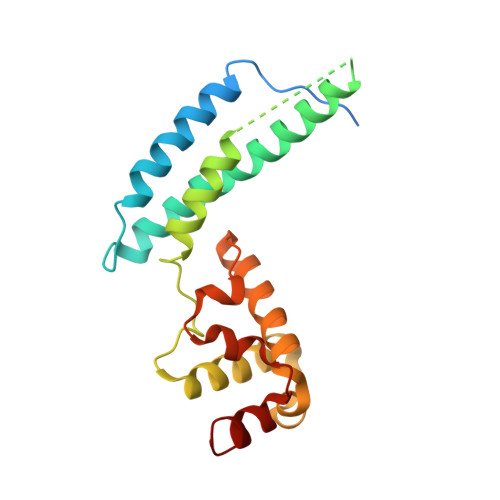
RNA Silencing Suppressor p21 of Beet Yellows Virus Forms an RNA Binding Octameric Ring Structure
Ye, K., Patel, D.J.(2005) Structure 13: 1375-1384
- PubMed: 16154094 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2005.06.017
- Primary Citation Related Structures:
2CWO - PubMed Abstract:
Many plant viruses encode proteins that suppress the antiviral RNA silencing response mounted by the host. The suppressors p19 from tombusvirus and p21 from Beet yellows virus appear to block silencing by directly binding siRNA, a critical mediator in the process. Here, we report the crystal structure of p21, which reveals an octameric ring architecture with a large central cavity of approximately 90 A diameter. The all alpha-helical p21 monomer consists of N- and C-terminal domains that associate with their neighboring counterparts through symmetric head-to-head and tail-to-tail interactions. A putative RNA binding surface is identified in the conserved, positive-charged inner surface of the ring. In contrast to the specific p19-siRNA duplex interaction, p21 is a general nucleic acid binding protein, interacting with 21 nt or longer single- and double-stranded RNAs in vitro. This study reveals an RNA binding structure adopted by the p21 silencing suppressor.
- Structural Biology Program, Memorial Sloan-Kettering Cancer Center, 1275 York Avenue, New York, NY 10021, USA.
Organizational Affiliation: