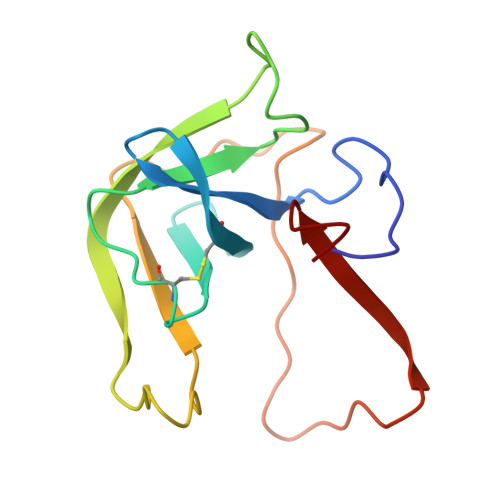
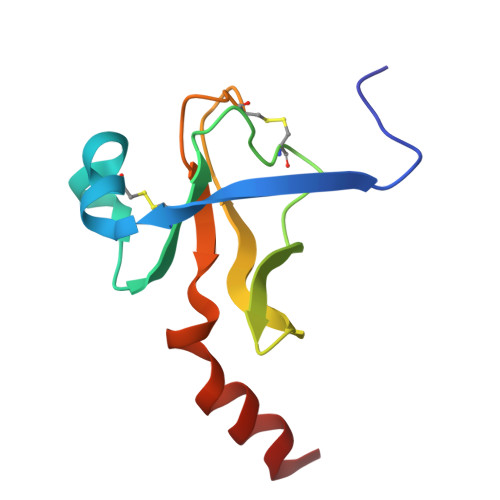
Structure of crystalline -chymotrypsin. V. The atomic structure of tosyl- -chymotrypsin at 2 A resolution.
Birktoft, J.J., Blow, D.M.(1972) J Mol Biology 68: 187-240
- PubMed: 5069789 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(72)90210-0
- Primary Citation Related Structures:
2CHA