Controlling Quaternary Structure Assembly: Subunit Interface Engineering and Crystal Structure of Dual Chain Avidin.
Hytonen, V.P., Horha, J., Airenne, T.T., Niskanen, E.A., Helttunen, K., Johnson, M.S., Salminen, T.A., Kulomaa, M.S., Nordlund, H.R.(2006) J Mol Biology 359: 1352
- PubMed: 16787776 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.04.044
- Primary Citation Related Structures:
2C4I - PubMed Abstract:
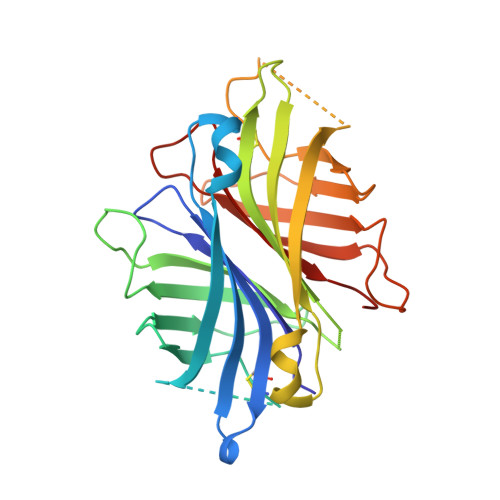
Dual chain avidin (dcAvd) is an engineered avidin form, in which two circularly permuted chicken avidin monomers are fused into one polypeptide chain. DcAvd can theoretically form two different pseudotetrameric quaternary assemblies because of symmetry at the monomer-monomer interfaces. Here, our aim was to control the assembly of the quaternary structure of dcAvd. We introduced the mutation I117C into one of the circularly permuted domains of dcAvd and scanned residues along the 1-3 subunit interface of the other domain. Interestingly, V115H resulted in a single, disulfide locked quaternary assembly of dcAvd, whereas I117H could not guide the oligomerisation process even though it stabilised the protein. The modified dcAvd forms were found to retain their characteristic pseudotetrameric state both at high and low pH, and were shown to bind D-biotin at levels comparable to that of wild-type chicken avidin. The crystal structure of dcAvd-biotin complex at 1.95 Angstroms resolution demonstrates the formation of the functional dcAvd pseudotetramer at the atomic level and reveals the molecular basis for its special properties. Altogether, our data facilitate further engineering of the biotechnologically valuable dcAvd scaffold and gives insights into how to guide the quaternary structure assembly of oligomeric proteins.
- NanoScience Center, Department of Biological and Environmental Science, University of Jyväskylä, Finland.
Organizational Affiliation: