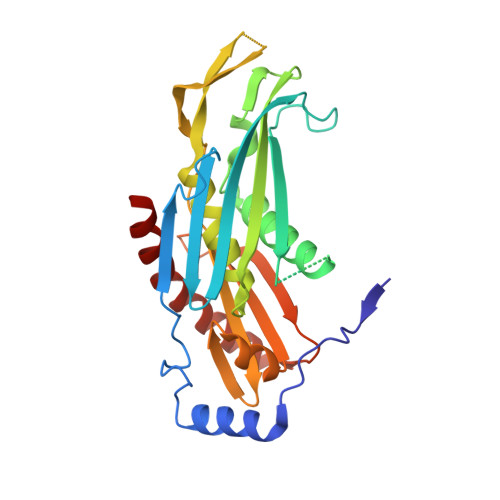
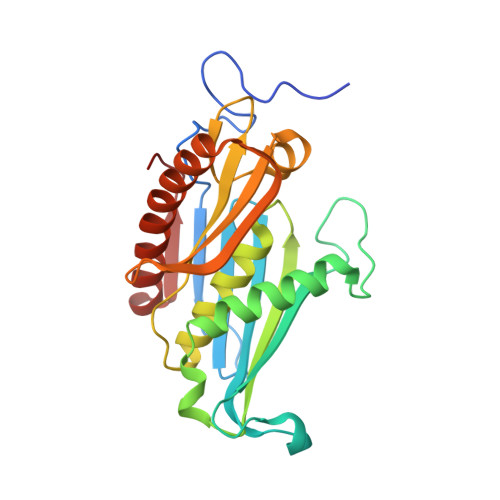
Structural Basis of 3' End RNA Recognition and Exoribonucleolytic Cleavage by an Exosome Rnase Ph Core.
Lorentzen, E., Conti, E.(2005) Mol Cell 20: 473
- PubMed: 16285928 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2005.10.020
- Primary Citation Related Structures:
2C37, 2C38, 2C39 - PubMed Abstract:
The exosome is a macromolecular complex that plays fundamental roles in the biogenesis and turnover of a large number of RNA species. Here we report the crystal structures of the Rrp41-Rrp42 core complex of the S. solfataricus exosome bound to short single-stranded RNAs and to ADP. The RNA binding cleft recognizes four nucleotides in a sequence-unspecific manner, mainly by electrostatic interactions with the phosphate groups. Interactions at the 2' hydroxyls of the sugars provide specificity for RNA over DNA. The structures show both the bound substrate and the cleaved product of the reaction, suggesting a catalytic mechanism for the 3'-5' phosphorolytic activity of the exosome.
- European Molecular Biology Laboratory, Meyerhofstrasse 1, D-69117 Heidelberg, Germany.
Organizational Affiliation: