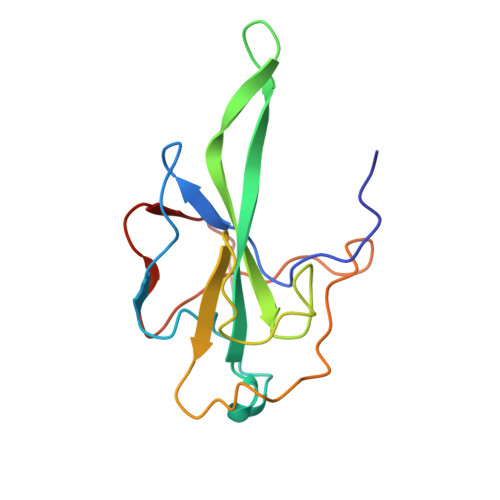
The Nucleocapsid Protein of Coronavirus Infectious Bronchitis Virus: Crystal Structure of its N-Terminal Domain and Multimerization Properties.
Fan, H., Ooi, A., Tan, Y.W., Wang, S., Fang, S., Liu, D.-X., Lescar, J.(2005) Structure 13: 1859
- PubMed: 16338414 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2005.08.021
- Primary Citation Related Structures:
2BTL, 2BXX - PubMed Abstract:
The coronavirus nucleocapsid (N) protein packages viral genomic RNA into a ribonucleoprotein complex. Interactions between N proteins and RNA are thus crucial for the assembly of infectious virus particles. The 45 kDa recombinant nucleocapsid N protein of coronavirus infectious bronchitis virus (IBV) is highly sensitive to proteolysis. We obtained a stable fragment of 14.7 kDa spanning its N-terminal residues 29-160 (IBV-N29-160). Like the N-terminal RNA binding domain (SARS-N45-181) of the severe acute respiratory syndrome virus (SARS-CoV) N protein, the crystal structure of the IBV-N29-160 fragment at 1.85 A resolution reveals a protein core composed of a five-stranded antiparallel beta sheet with a positively charged beta hairpin extension and a hydrophobic platform that are probably involved in RNA binding. Crosslinking studies demonstrate the formation of dimers, tetramers, and higher multimers of IBV-N. A model for coronavirus shell formation is proposed in which dimerization of the C-terminal domain of IBV-N leads to oligomerization of the IBV-nucleocapsid protein and viral RNA condensation.
- School of Biological Sciences, Nanyang Technological University, 60 Nanyang Drive, Singapore 637551.
Organizational Affiliation: