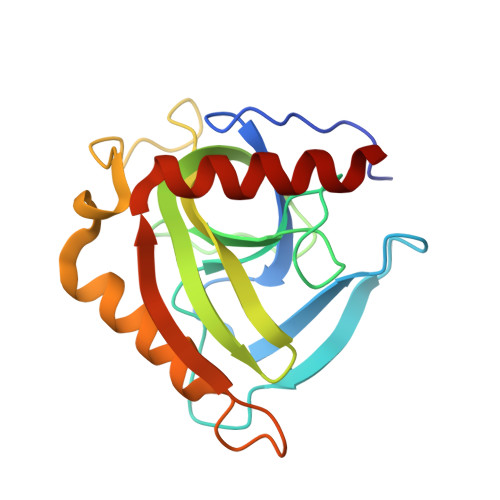
Kinetic and Structural Properties of Inorganic Pyrophosphatase from the Pathogenic Bacterium Helicobacter Pylori.
Chao, T.-C., Huang, H., Tsai, J.Y., Huang, C.Y., Sun, Y.-J.(2006) Proteins 65: 670
- PubMed: 16988955 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21093
- Primary Citation Related Structures:
2BQX, 2BQY - PubMed Abstract:
Inorganic pyrophosphatase (PPase) catalyzes the hydrolysis of pyrophosphate (PPi) to orthophosphate (Pi) and controls the level of PPi in cells. PPase plays an essential role in energy conservation and provides the energy for many biosynthetic pathways. The Helicobacter pylori pyrophosphatase (HpPPase) gene was cloned, expressed, purified, and found to have a molecular weight of 20 kDa. The K(m) and V (max) of HpPPase were determined as 214.4 microM and 594 micromol Pi min(-1) mg(-1), respectively. PPi binds Mg(2+) to form a true substrate that activates the enzyme. However, free PPi could be a potent inhibitor for HpPPase. The effects of the inhibitors NaF, ATP, iminodiphosphate, and N-ethylmaleimide on HpPPase activity were evaluated. NaF showed the highest inhibition of the enzyme. Crystal structures of HpPPase and the PPi-HpPPase complex were determined. HpPPase comprises three alpha-helices and nine beta-strands and folds as a barrel structure. HpPPase forms a hexamer in both the solution and crystal states, and each monomer has its own PPi-binding site. The PPi binding does not cause a significant conformational change in the PPi-HpPPase complex, which might represent an inhibition state for HpPPase in the absence of a divalent metal ion.
- Institute of Bioinformatics and Structural Biology, National Tsing Hua University, Hsinchu 300, Taiwan, Republic of China.
Organizational Affiliation: