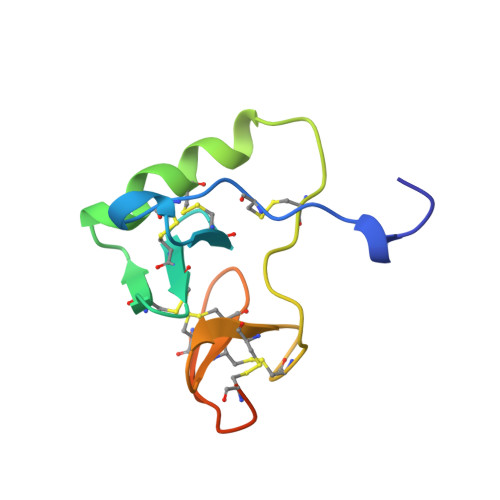
Crystal structure of a bovine neurophysin II dipeptide complex at 2.8 A determined from the single-wavelength anomalous scattering signal of an incorporated iodine atom.
Chen, L.Q., Rose, J.P., Breslow, E., Yang, D., Chang, W.R., Furey Jr., W.F., Sax, M., Wang, B.C.(1991) Proc Natl Acad Sci U S A 88: 4240-4244
- PubMed: 2034668 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.88.10.4240
- Primary Citation Related Structures:
2BN2 - PubMed Abstract:
The crystal structure of a dipeptide complex of bovine neurophysin II has been solved at 2.8 A resolution solely by using single-wavelength anomalous scattering data from a single iodinated derivative. The asymmetric unit is an elongated tetramer of dimensions 110 x 40 x 30 A, composed of two dimers related by pseudo twofold symmetry. Each monomer consists of two homologous layers, each with four antiparallel beta-strands. The two regions are connected by a helix followed by a long loop. Monomer-monomer contacts involve antiparallel beta-sheet interactions, which form a dimer with two layers of eight beta-strands. One peptide per monomer occupies the principal hormone-binding pocket formed by part of the amino-terminal region and parts of the connecting helix and loop, with binding to protein consistent with conclusions drawn from solution studies. Dimer-dimer contacts involve the Tyr49 region adjacent to this site. A fifth dipeptide, of unknown biological significance, helps to stabilize one of the monomer-monomer interfaces and the tetramer-tetramer network in the crystal.
- Department of Crystallography, University of Pittsburgh, PA 15260.
Organizational Affiliation: