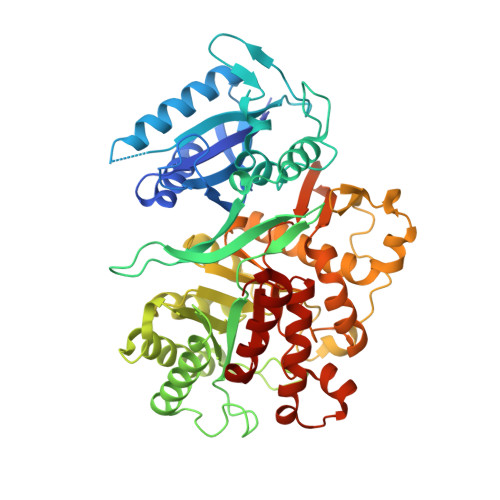
The Three-Dimensional Structure of the Bifunctional 6-Hydroxymethyl-7,8-Dihydropterin Pyrophosphokinase/Dihydropteroate Synthase of Saccharomyces Cerevisiae
Lawrence, M.C., Iliades, P., Fernley, R.T., Berglez, J., Pilling, P.A., Macreadie, I.G.(2005) J Mol Biology 348: 655
- PubMed: 15826662 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.03.021
- Primary Citation Related Structures:
2BMB - PubMed Abstract:
In Saccharomyces cerevisiae and other fungi, the enzymes dihydroneopterin aldolase, 6-hydroxymethyl-7,8-dihydropterin pyrophosphokinase (HPPK) and dihydropteroate synthase (DHPS) are encoded by a polycistronic gene that is translated into a single polypeptide having all three functions. These enzymatic functions are essential to both prokaryotes and lower eukaryotes, and catalyse sequential reactions in folate biosynthesis. Deletion or disruption of either function leads to cell death. These enzymes are absent from mammals and thus make ideal antimicrobial targets. DHPS is currently the target of antifolate therapy for a number of infectious diseases, and its activity is inhibited by sulfonamides and sulfones. These drugs are typically used as part of a synergistic cocktail with the 2,4-diaminopyrimidines that inhibit dihydrofolate reductase. A gene encoding the S.cerevisiae HPPK and DHPS enzymes has been cloned and expressed in Escherichia coli. A complex of the purified bifunctional polypeptide with a pterin monophosphate substrate analogue has been crystallized, and its structure solved by molecular replacement and refined to 2.3A resolution. The polypeptide consists of two structural domains, each of which closely resembles its respective monofunctional bacterial HPPK and DHPS counterpart. The mode of ligand binding is similar to that observed in the bacterial enzymes. The association between the domains within the polypeptide as well as the quaternary association of the polypeptide via its constituent DHPS domains provide insight into the assembly of the trifunctional enzyme in S.cerevisiae and probably other fungal species.
- CSIRO Health Sciences and Nutrition, 343 Royal Parade, Parkville, Victoria 3052, Australia. mike.lawrence@csiro.au
Organizational Affiliation: