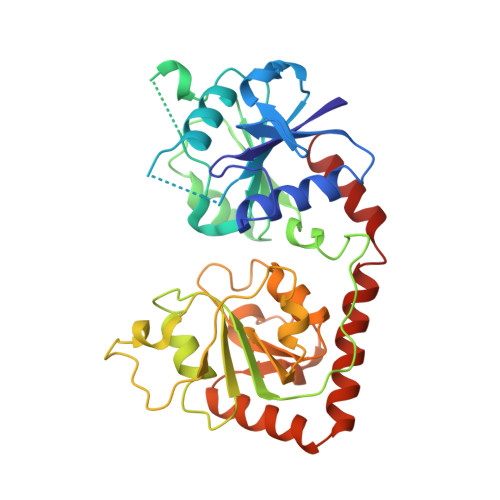
Crystal structure of the DNA modifying enzyme beta-glucosyltransferase in the presence and absence of the substrate uridine diphosphoglucose.
Vrielink, A., Ruger, W., Driessen, H.P., Freemont, P.S.(1994) EMBO J 13: 3413-3422
- PubMed: 8062817 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/j.1460-2075.1994.tb06646.x
- Primary Citation Related Structures:
1BGT, 1BGU, 2BGT, 2BGU - PubMed Abstract:
Bacteriophage T4 beta-glucosyltransferase (EC 2.4.1.27) catalyses the transfer of glucose from uridine diphosphoglucose to hydroxymethyl groups of modified cytosine bases in T4 duplex DNA forming beta-glycosidic linkages. The enzyme forms part of a phage DNA protection system. We have solved and refined the crystal structure of recombinant beta-glucosyltransferase to 2.2 A resolution in the presence and absence of the substrate, uridine diphosphoglucose. The structure comprises two domains of similar topology, each reminiscent of a nucleotide binding fold. The two domains are separated by a central cleft which generates a concave surface along one side of the molecule. The substrate-bound complex reveals only clear electron density for the uridine diphosphate portion of the substrate. The UDPG is bound in a pocket at the bottom of the cleft between the two domains and makes extensive hydrogen bonding contacts with residues of the C-terminal domain only. The domains undergo a rigid body conformational change causing the structure to adopt a more closed conformation upon ligand binding. The movement of the domains is facilitated by a hinge region between residues 166 and 172. Electrostatic surface potential calculations reveal a large positive potential along the concave surface of the structure, suggesting a possible site for duplex DNA interaction.
- Protein Structure Laboratory, Imperial Cancer Research Fund, London.
Organizational Affiliation: