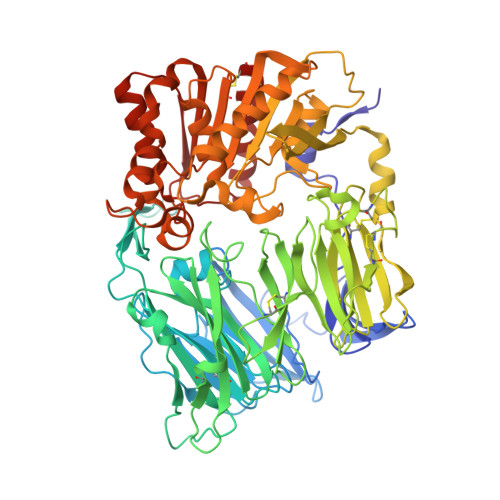
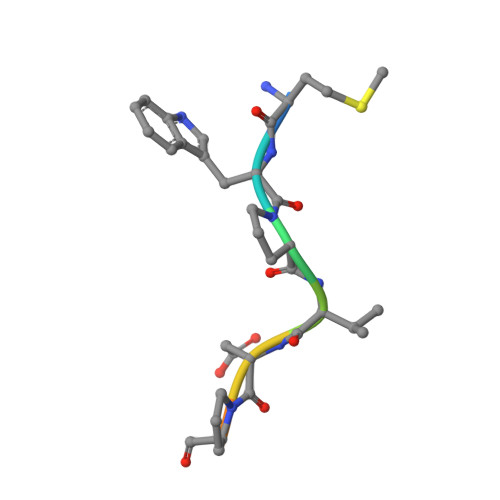
Crystal Structures of HIV-1 Tat-Derived Nonapeptides Tat-(1-9) and Trp2-Tat-(1-9) Bound to the Active Site of Dipeptidyl-Peptidase Iv (Cd26)
Weihofen, W.A., Liu, J., Reutter, W., Saenger, W., Fan, H.(2005) J Biological Chem 280: 14911
- PubMed: 15695814 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M413400200
- Primary Citation Related Structures:
2BGN, 2BGR - PubMed Abstract:
CD26 or dipeptidyl-peptidase IV (DPPIV) is engaged in immune functions by co-stimulatory effects on activation and proliferation of T lymphocytes, binding to adenosine deaminase, and regulation of various chemokines and cytokines. DPPIV peptidase activity is inhibited by both Tat protein from human immunodeficiency virus (HIV)-1 and its N-terminal nonapeptide Tat-(1-9) with amino acid sequence MDPVDPNIE, suggesting that DPPIV mediates immunosuppressive effects of Tat protein. The 2.0- and 3.15-A resolution crystal structures of the binary complex between human DPPIV and nonapeptide Tat-(1-9) and the ternary complex between the variant MWPVDPNIE, called Trp(2)-Tat-(1-9), and DPPIV bound to adenosine deaminase show that Tat-(1-9) and Trp(2)-Tat-(1-9) are located in the active site of DPPIV. The interaction pattern of DPPIV with Trp(2)-Tat-(1-9) is tighter than that with Tat-(1-9), in agreement with inhibition constants (K(i)) of 2 x 10(-6) and 250 x 10(-6) m, respectively. Both peptides cannot be cleaved by DPPIV because the binding pockets of the N-terminal 2 residues are interchanged compared with natural substrates: the N-terminal methionine occupies the hydrophobic S1 pocket of DPPIV that normally accounts for substrate specificity by binding the penultimate residue. Because the N-terminal sequence of the thromboxane A2 receptor resembles the Trp(2)-Tat-(1-9) peptide, a possible interaction with DPPIV is postulated.
- Institut für Chemie/Kristallographie, Freie Universität Berlin, Takustrasse 6, D-14195 Berlin, Germany.
Organizational Affiliation: