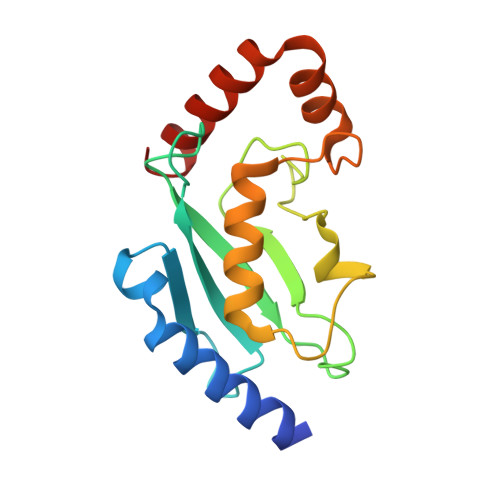
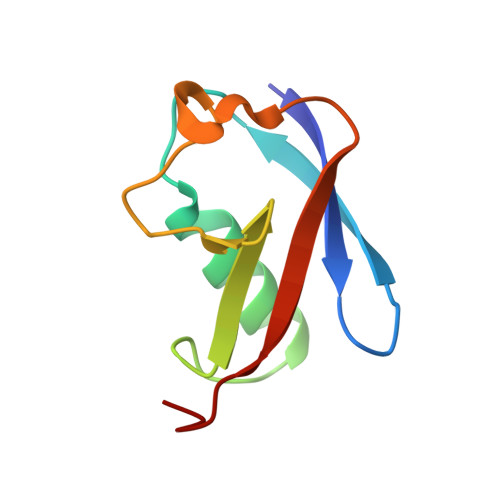
Sumo Imodification of the Ubiquitin Conjugating Enzyme E2-25K
Pichler, A., Knipscheer, P., Oberhofer, E., Van Dijk, W.J., Korner, R., Velgaard Olsen, J., Jentsch, S., Melchior, F., Sixma, T.K.(2005) Nat Struct Mol Biol 12: 264
- PubMed: 15723079 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb903
- Primary Citation Related Structures:
2BEP, 2BF8 - PubMed Abstract:
Post-translational modification with small ubiquitin-related modifier (SUMO) alters the function of many proteins, but the molecular mechanisms and consequences of this modification are still poorly defined. During a screen for novel SUMO1 targets, we identified the ubiquitin-conjugating enzyme E2-25K (Hip2). SUMO attachment severely impairs E2-25K ubiquitin thioester and unanchored ubiquitin chain formation in vitro. Crystal structures of E2-25K(1-155) and of the E2-25K(1-155)-SUMO conjugate (E2-25K(*)SUMO) indicate that SUMO attachment interferes with E1 interaction through its location on the N-terminal helix. The SUMO acceptor site in E2-25K, Lys14, does not conform to the consensus site found in most SUMO targets (PsiKXE), and functions only in the context of an alpha-helix. In contrast, adjacent SUMO consensus sites are modified only when in unstructured peptides. The demonstration that secondary structure elements are part of SUMO attachment signals could contribute to a better prediction of SUMO targets.
- University of Göttingen, Department of Biochemistry I, Humboldt Allee 23, 37073 Göttingen, Germany.
Organizational Affiliation: