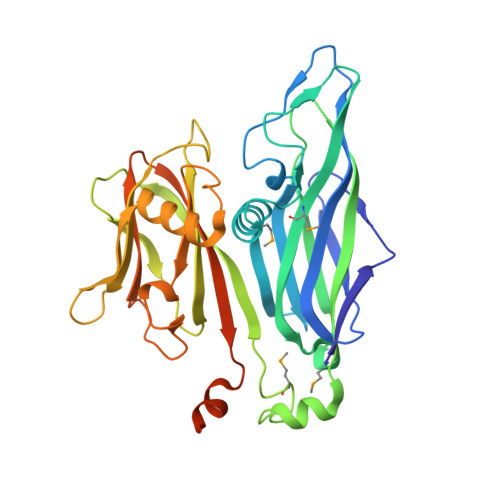
Structure of an archaeal virus capsid protein reveals a common ancestry to eukaryotic and bacterial viruses.
Khayat, R., Tang, L., Larson, E.T., Lawrence, C.M., Young, M., Johnson, J.E.(2005) Proc Natl Acad Sci U S A 102: 18944-18949
- PubMed: 16357204 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0506383102
- Primary Citation Related Structures:
2BBD - PubMed Abstract:
Archaea and their viruses are poorly understood when compared with the Eukarya and Bacteria domains of life. We report here the crystal structure of the major capsid protein (MCP) of the Sulfolobus turreted icosahedral virus, an archaeal virus isolated from an acidic hot spring (pH 2-4, 72-92 degrees C) in Yellowstone National Park. The structure is nearly identical to the MCP structures of the eukaryotic Paramecium bursaria Chlorella virus, and the bacteriophage PRD1, and shows a common fold with the mammalian adenovirus. Structural analysis of the capsid architecture, determined by fitting the subunit into the electron cryomicroscopy reconstruction of the virus, identified a number of key interactions that are akin to those observed in adenovirus and PRD1. The similar capsid proteins and capsid architectures strongly suggest that these viral capsids originated and evolved from a common ancestor. Hence, this work provides a previously undescribed example of a viral relationship spanning the three domains of life (Eukarya, Bacteria, and Archaea). The MCP structure also provides insights into the stabilizing forces required for extracellular hyperthermophilic proteins to tolerate high-temperature hot springs.
- Department of Molecular Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: