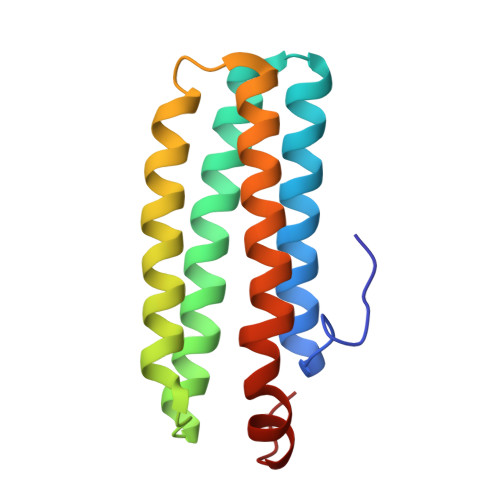
Structural Basis for O(2) Sensing by the Hemerythrin-like Domain of a Bacterial Chemotaxis Protein: Substrate Tunnel and Fluxional N Terminus
Isaza, C.E., Silaghi-Dumitrescu, R., Iyer, R.B., Kurtz, D.M., Chan, M.K.(2006) Biochemistry 45: 9023-9031
- PubMed: 16866347 Search on PubMed
- DOI: https://doi.org/10.1021/bi0607812
- Primary Citation Related Structures:
2AVK, 2AWC, 2AWY - PubMed Abstract:
The methyl-accepting chemotaxis protein, DcrH, from the anaerobic sulfate-reducing bacterium, Desulfovibrio vulgaris (Hildenborough), has a hemerythrin-like domain, DcrH-Hr, at its C terminus. DcrH-Hr was previously shown to contain a diiron site that binds O2, suggesting an O2-sensing function. X-ray crystal structures of diferric (met-), azido-diferric (azidomet-), and diferrous (deoxy-) DcrH-Hr reveal a "substrate tunnel" distinct from that in invertebrate hemerythrins. This tunnel is proposed to facilitate the rapid autoxidation of oxy-DcrH-Hr and suggests that sensing is triggered by O2 binding and subsequent oxidation of the diferrous active site. The N-terminal loop of DcrH-Hr is highly ordered in both met- and azidomet-DcrH-Hr but is disordered in deoxy-DcrH-Hr. These redox-dependent conformational differences presumably transduce the sensory signal of DcrH-Hr to the neighboring methylation domain in the full-length receptor. Given the putative cytoplasmic localization of its Hr-like O2-sensing domain, DcrH is proposed to serve a role in negative aerotaxis (anaerotaxis).
- The Ohio State Biophysics Program, The Ohio State University, 484 West 12th Avenue, Columbus, Ohio 43210, USA.
Organizational Affiliation: