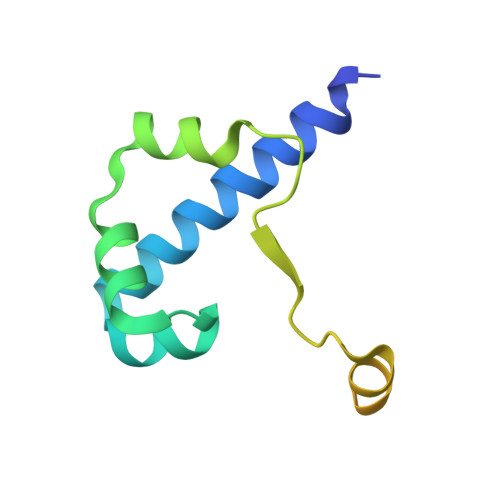
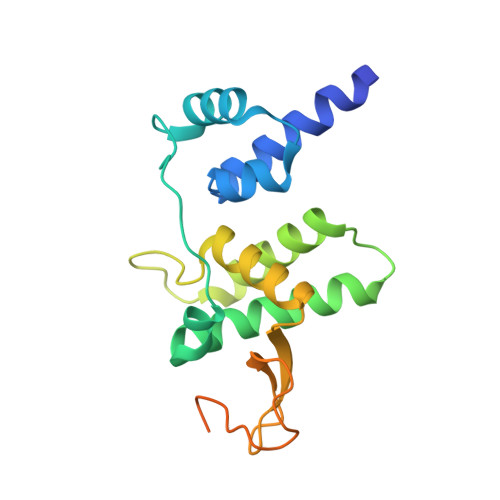
Structure of the Escherichia coli FlhDC Complex, a Prokaryotic Heteromeric Regulator of Transcription.
Wang, S., Fleming, R.T., Westbrook, E.M., Matsumura, P., McKay, D.B.(2006) J Mol Biology 355: 798-808
- PubMed: 16337229 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.11.020
- Primary Citation Related Structures:
2AVU - PubMed Abstract:
The hetero-oligomeric complex of the FlhD and FlhC proteins (FlhDC) regulates transcription from several flagellar and non-flagellar operons in bacteria. The crystallographic structure of the Escherichia coli FlhDC complex has been solved to 3.0 A resolution, revealing a hexameric FlhD4FlhC2 assembly. In the complex, each FlhC protomer binds an FlhD2 dimer; the conformation of the dimer in the complex differs significantly from its conformation in the absence of FlhC. FlhC has a novel tertiary fold that includes a heretofore unrecognized zinc-binding site in which the ion is ligated by four cysteine residues. Gel shift experiments show that binding of the FlhDC complex to a cognate promoter bends the DNA by approximately 111 degrees . The structure of the FlhDC complex is compatible with models in which a fragment of operator DNA, at least 48 base-pairs in length, wraps around the complex and bends significantly when binding.
- Department of Structural Biology, Stanford University School of Medicine, Stanford, CA 94305, USA.
Organizational Affiliation: