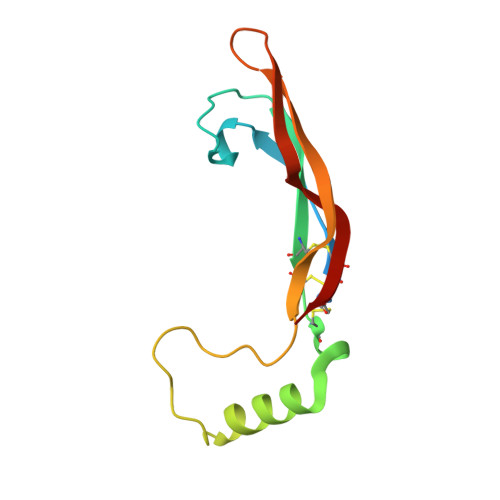
Artemin crystal structure reveals insights into heparan sulfate binding.
Silvian, L., Jin, P., Carmillo, P., Boriack-Sjodin, P.A., Pelletier, C., Rushe, M., Gong, B.J., Sah, D., Pepinsky, B., Rossomando, A.(2006) Biochemistry 45: 6801-6812
- PubMed: 16734417 Search on PubMed
- DOI: https://doi.org/10.1021/bi060035x
- Primary Citation Related Structures:
2ASK - PubMed Abstract:
Artemin (ART) promotes the growth of developing peripheral neurons by signaling through a multicomponent receptor complex comprised of a transmembrane tyrosine kinase receptor (cRET) and a specific glycosylphosphatidylinositol-linked co-receptor (GFRalpha3). Glial cell line-derived neurotrophic factor (GDNF) signals through a similar ternary complex but requires heparan sulfate proteoglycans (HSPGs) for full activity. HSPG has not been demonstrated as a requirement for ART signaling. We crystallized ART in the presence of sulfate and solved its structure by isomorphous replacement. The structure reveals ordered sulfate anions bound to arginine residues in the pre-helix and amino-terminal regions that were organized in a triad arrangement characteristic of heparan sulfate. Three residues in the pre-helix were singly or triply substituted with glutamic acid, and the resulting proteins were shown to have reduced heparin-binding affinity that is partly reflected in their ability to activate cRET. This study suggests that ART binds HSPGs and identifies residues that may be involved in HSPG binding.
- Department of Drug Discovery, Biogen Idec, Inc., 12 Cambridge Center, Cambridge, Massachusetts 02142, USA. laura.silvian@biogenidec.com
Organizational Affiliation: