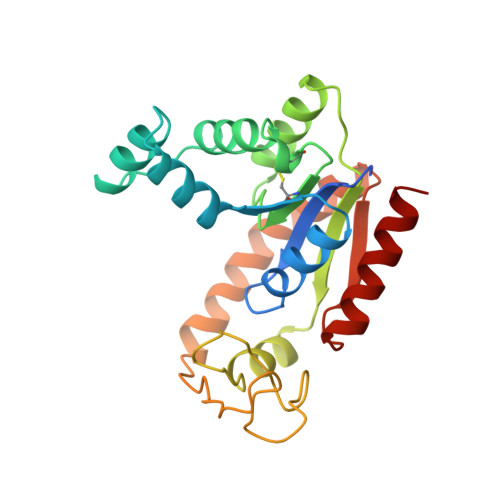
The structure of bovine mitochondrial adenylate kinase: comparison with isoenzymes in other compartments.
Schlauderer, G.J., Schulz, G.E.(1996) Protein Sci 5: 434-441
- PubMed: 8868479 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.5560050304
- Primary Citation Related Structures:
1AK2, 2AK2 - PubMed Abstract:
In vertebrates, there are different adenylate kinases in the compartments cytosol, mitochondrial intermembrane space, and mitochondrial matrix. Here, we report the spatial structure of the intermembrane species established in two crystal forms by X-ray diffraction analyses at 1.92 and 2.1 A resolution. In both structures, the enzyme is unligated, and thus in an "open" conformation. The enzyme was prepared from bovine liver, containing at least five variants arisen from posttranscriptional and posttranslational modifications. It could only be crystallized after removing some of these variants. A comparison with the known structures of the adenylate kinases from cytosol and mitochondrial matrix reveals structural differences that should play a role in protein targeting because none of these enzymes contains a cleavable signal peptide. A further comparison with adenylate kinases from Gram-positive bacteria showed that the structural Zn2+ ion of these species is replaced by a strictly conserved assembly of hydrogen bonded residues.
- Institut für Organische Chemie und Biochemie, Freiburg im Breisgau, Germany.
Organizational Affiliation: