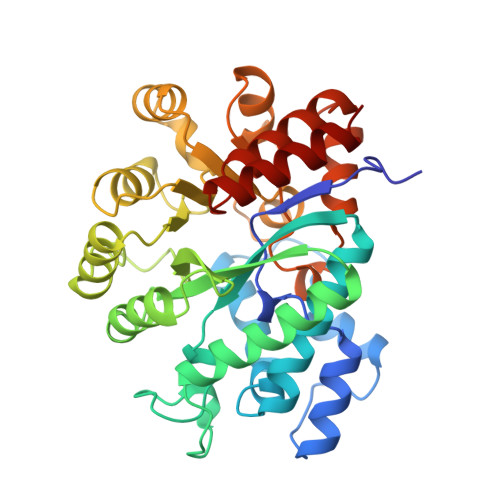
Atomic structure of adenosine deaminase complexed with a transition-state analog: understanding catalysis and immunodeficiency mutations.
Wilson, D.K., Rudolph, F.B., Quiocho, F.A.(1991) Science 252: 1278-1284
- PubMed: 1925539 Search on PubMed
- DOI: https://doi.org/10.1126/science.1925539
- Primary Citation Related Structures:
2ADA - PubMed Abstract:
The crystal structure of a murine adenosine deaminase complexed with 6-hydroxyl-1,6-dihydropurine ribonucleoside, a nearly ideal transition-state analog, has been determined and refined at 2.4 angstrom resolution. The structure is folded as an eight-stranded parallel alpha/beta barrel with a deep pocket at the beta-barrel COOH-terminal end wherein the inhibitor and a zinc are bound and completely sequestered. The presence of the zinc cofactor and the precise structure of the bound analog were not previously known. The 6R isomer of the analog is very tightly held in place by the coordination of the 6-hydroxyl to the zinc and the formation of nine hydrogen bonds. On the basis of the structure of the complex a stereoselective addition-elimination or SN2 mechanism of the enzyme is proposed with the zinc atom and the Glu and Asp residues playing key roles. A molecular explanation of a hereditary disease caused by several point mutations of an enzyme is also presented.
- Howard Hughes Medical Institute, Baylor College of Medicine, Houston, TX 77030.
Organizational Affiliation: