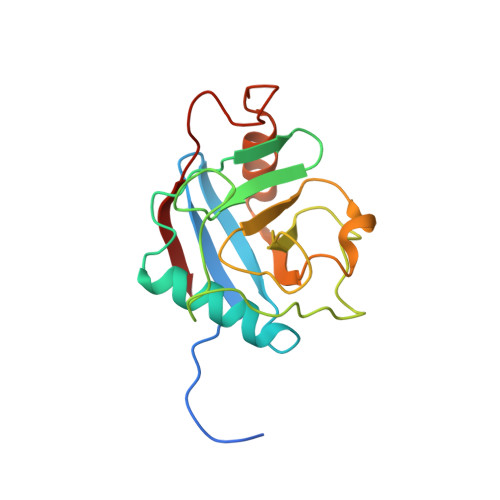
The crystal structure of human WD40 repeat-containing peptidylprolyl isomerase (PPWD1).
Davis, T.L., Walker, J.R., Ouyang, H., MacKenzie, F., Butler-Cole, C., Newman, E.M., Eisenmesser, E.Z., Dhe-Paganon, S.(2008) FEBS J 275: 2283-2295
- PubMed: 18397323 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2008.06381.x
- Primary Citation Related Structures:
2A2N - PubMed Abstract:
Cyclophilins comprise one of the three classes of peptidylprolyl isomerases found in all eukaryotic and prokaryotic organisms, as well as viruses. Many of the 17 annotated human cyclophilins contain the catalytic domain in tandem with other domains, and many of the specific functions of a particular cyclophilin or its associated domains remain unknown. The structure of the isomerase domain from a spliceosome-associated cyclophilin, PPWD1 (peptidylprolyl isomerase containing WD40 repeat), has been solved to 1.65 A. In the crystal, the N-terminus of one isomerase domain is bound in the active site of a neighboring isomerase molecule in a manner analogous to substrate. NMR solution studies show that this sequence binds to the active site of the cyclophilin, but cannot be turned over by the enzyme. A pseudo-substrate immediately N-terminal to the cyclophilin domain in PPWD1 could have wider implications for the function of this cyclophilin in the spliceosome, where it is located in human cells.
- Structural Genomics Consortium, Banting Institute, University of Toronto, ON, Canada.
Organizational Affiliation: