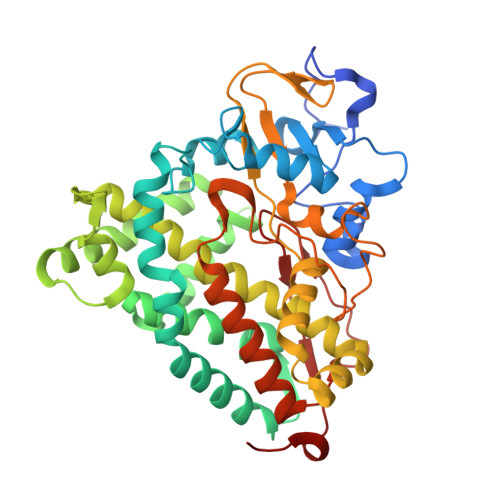
Crystallographic study on the dioxygen complex of wild-type and mutant cytochrome P450cam. Implications for the dioxygen activation mechanism
Nagano, S., Poulos, T.L.(2005) J Biological Chem 280: 31659-31663
- PubMed: 15994329 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M505261200
- Primary Citation Related Structures:
2A1M, 2A1N, 2A1O - PubMed Abstract:
Two key amino acids, Thr252 and Asp251, are known to be important for dioxygen activation by cytochrome P450cam. We have solved crystal structures of a critical intermediate, the ferrous dioxygen complex (Fe(II)-O2), of the wild-type P450cam and its mutants, D251N and T252A. The wild-type dioxygen complex structure is very much the same as reported previously (Schlichting, I., Berendzen, J., Chu, K., Stock, A. M., Maves, S. A., Benson, D. E., Sweet, R. M., Ringe, D., Petsko, G. A., and Sligar, S. G. (2000) Science 287, 1615-1622) with the exception of higher occupancy and a more ordered structure of the iron-linked dioxygen and two "catalytic" water molecules that form part of a proton relay system to the iron-linked dioxygen. Due to of the altered conformation of the I helix groove these two waters are missing in the D251N dioxygen complex which explains its lower catalytic activity and slower proton transfer to the dioxygen ligand. Similarly, the T252A mutation was expected to disrupt the active site solvent structure leading to hydrogen peroxide formation rather than substrate hydroxylation. Unexpectedly, however, the two "catalytic" waters are retained in the T252A mutant. Based on these findings, we propose that the Thr(252) accepts a hydrogen bond from the hydroperoxy (Fe(III)-OOH) intermediate that promotes the second protonation on the distal oxygen atom, leading to O-O bond cleavage and compound I formation.
- Department of Molecular Biology & Biochemistry, University of California, Irvine 92697-3900, USA. snagano@riken.jp
Organizational Affiliation: